ACMG/AMP interpretation of BRCA1 missense variants: Structure-informed scores add evidence strength granularity to the PP3/BP4 computational evidence
- PMID: 40233743
- PMCID: PMC12120176
- DOI: 10.1016/j.ajhg.2024.12.011
ACMG/AMP interpretation of BRCA1 missense variants: Structure-informed scores add evidence strength granularity to the PP3/BP4 computational evidence
Abstract
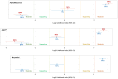
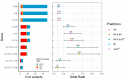
Classification of missense variants is challenging. Lacking compelling clinical and/or functional data, ACMG/AMP lines of evidence are restricted to PM2 (rarity code applied at supporting level) and PP3/BP4 (computational evidence based mostly on multiple-sequence-alignment conservation tools). Currently, the ClinGen ENIGMA BRCA1/2 Variant Curation Expert Panel uses BayesDel to apply PP3/BP4 to missense variants located in the BRCA1 RING/BRCT domains. The ACMG/AMP framework does not refer explicitly to protein structure as a putative source of pathogenic/benign evidence. Here, we tested the value of incorporating structure-based evidence such as relative solvent accessibility (RSA), folding stability (ΔΔG), and/or AlphaMissense pathogenicity to the classification of BRCA1 missense variants. We used MAVE functional scores as proxies for pathogenicity/benignity. We computed RSA and FoldX5.0 ΔΔG predictions using as alternative input templates for either PDB files or AlphaFold2 models, and we retrieved pre-computed AlphaMissense and BayesDel scores. We calculated likelihood ratios toward pathogenicity/benignity provided by the tools (individually or combined). We performed a clinical validation of major findings using the large-scale BRIDGES case-control dataset. AlphaMissense outperforms ΔΔG and BayesDel, providing similar PP3/BP4 evidence strengths with lower rate of variants in the uninformative score range. AlphaMissense combined with ΔΔG increases evidence strength granularity. AlphaFold2 models perform well as input templates for ΔΔG predictions. Regardless of the tool, BP4 (but not PP3) is highly dependent on RSA, with benignity evidence provided only to variants targeting buried or partially buried residues (RSA ≤ 60%). Stratification by functional domain did not reveal major differences. In brief, structure-based analysis improves PP3/BP4 assessment, uncovering a relevant role for RSA.
Keywords: ACMG/AMP; AlphaFold; AlphaMissense; BRCA1; BayesDel; MAVE; PP3/BP4; RSA; ΔΔG.
Copyright © 2024 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests M.J.V. is an employee of Ambry Genetics. A.C. is an employee of Ambry Genetics. M.E.R. is an employee of Ambry Genetics.
Figures



Similar articles
-
Structural biology in variant interpretation: Perspectives and practices from two studies.Am J Hum Genet. 2025 May 1;112(5):984-992. doi: 10.1016/j.ajhg.2025.03.010. Epub 2025 Apr 14. Am J Hum Genet. 2025. PMID: 40233741 Free PMC article. Review.
-
Integration of protein stability and AlphaMissense scores improves bioinformatic impact prediction for p53 missense and in-frame amino acid deletion variants.Am J Hum Genet. 2025 May 1;112(5):1003-1014. doi: 10.1016/j.ajhg.2025.01.012. Epub 2025 Apr 14. Am J Hum Genet. 2025. PMID: 40233742
-
Assessment of the evidence yield for the calibrated PP3/BP4 computational recommendations.Genet Med. 2024 Nov;26(11):101213. doi: 10.1016/j.gim.2024.101213. Epub 2024 Jul 25. Genet Med. 2024. PMID: 39030733
-
Calibration of computational tools for missense variant pathogenicity classification and ClinGen recommendations for PP3/BP4 criteria.Am J Hum Genet. 2022 Dec 1;109(12):2163-2177. doi: 10.1016/j.ajhg.2022.10.013. Epub 2022 Nov 21. Am J Hum Genet. 2022. PMID: 36413997 Free PMC article.
-
DNA repair-related functional assays for the classification of BRCA1 and BRCA2 variants: a critical review and needs assessment.J Med Genet. 2017 Nov;54(11):721-731. doi: 10.1136/jmedgenet-2017-104707. Epub 2017 Sep 2. J Med Genet. 2017. PMID: 28866612 Review.
Cited by
-
Structural biology in variant interpretation: Perspectives and practices from two studies.Am J Hum Genet. 2025 May 1;112(5):984-992. doi: 10.1016/j.ajhg.2025.03.010. Epub 2025 Apr 14. Am J Hum Genet. 2025. PMID: 40233741 Free PMC article. Review.
References
-
- Schaafsma G.C.P., Vihinen M. Large differences in proportions of harmful and benign amino acid substitutions between proteins and diseases. Hum. Mutat. 2017;38:1613–1848. - PubMed
-
- Casadio R., Vassura M., Tiwari S., Fariselli P., Luigi Martelli P. Correlating disease-related mutations to their effect on protein stability: a large-scale analysis of the human proteome. Hum. Mutat. 2011;32:1161–1170. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

