This is a preprint.
Discovery and therapeutic exploitation of Master Regulatory miRNAs in Glioblastoma
- PMID: 40236125
- PMCID: PMC11996502
- DOI: 10.1101/2025.04.01.646663
Discovery and therapeutic exploitation of Master Regulatory miRNAs in Glioblastoma
Abstract
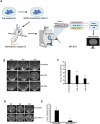
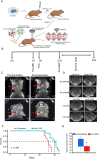
Glioblastoma is a fatal primary malignant brain tumor, with an average survival of only 15 months despite surgical resection, chemotherapy, and radiation therapy. Due to the concurrent deregulation of numerous genes in glioblastoma, molecular monotherapies have not improved clinical outcomes. Evidence suggests that effectively targeting multiple deregulated molecules is essential for better therapies; however, this is limited by the lack of suitable drugs and the increased toxicity of combination therapies. To address this, we hypothesized that miRNAs, small gene-regulatory RNAs that suppress multiple target genes via sequence complementarity, could be developed to inhibit multiple deregulated genes simultaneously, leading to more effective treatments. We identified master regulatory miRNAs-those that target several deregulated genes in glioblastoma-using PAR-CLIP screenings in glioblastoma cells and analyzed TCGA tumor data to find which targets were deregulated. An algorithm ranked these targets based on their significance in glioblastoma malignancy. We selected two tumor suppressor master regulatory miRNAs, miR-340 and miR-382, and one oncogenic miRNA, miR-17. Validation showed that these miRNAs target critical glioblastoma pathways and significantly inhibit cell growth, survival, invasion, and tumor growth in vivo. We developed an innovative therapeutic delivery approach using Brain Penetrating Nanoparticles in combination with MRI-guided focused ultrasound and microbubbles, resulting in reduced tumor volume and extended survival in glioblastoma-bearing mice. This strategy offers a promising pathway for translating miRNA-based therapies into clinical trials for glioblastoma and other cancers.
Conflict of interest statement
Competing interests: The authors declare there are no competing interests.
Figures






References
-
- Bellail A. C., Hunter S. B., Brat D. J., Tan C., Van Meir E. G., Microregional extracellular matrix heterogeneity in brain modulates glioma cell invasion. The international journal of biochemistry & cell biology 36, 1046–1069 (2004). - PubMed
-
- Lim M., Xia Y., Bettegowda C., Weller M., Current state of immunotherapy for glioblastoma. Nature reviews Clinical oncology 15, 422–442 (2018). - PubMed
-
- Stupp R., Dietrich P.-Y., Kraljevic S. O., Pica A., Maillard I., Maeder P., Meuli R., Janzer R., Pizzolato G., Miralbell R., Promising survival for patients with newly diagnosed glioblastoma multiforme treated with concomitant radiation plus temozolomide followed by adjuvant temozolomide. Journal of Clinical Oncology 20, 1375–1382 (2002). - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
