RNF5 inhibits HBV replication by mediating caspase-3-dependent degradation of core protein
- PMID: 40236486
- PMCID: PMC11996839
- DOI: 10.3389/fmicb.2025.1548061
RNF5 inhibits HBV replication by mediating caspase-3-dependent degradation of core protein
Abstract
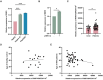
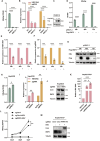
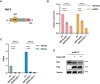
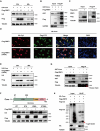
The RING finger protein 5 (RNF5), an E3 ubiquitin ligase, has demonstrated significant antiviral activity against various viruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and Kaposi's sarcoma-associated herpesvirus (KSHV). However, its role in hepatitis B virus (HBV) replication has not been previously studied. In this study, we demonstrate that RNF5 effectively inhibits HBV replication by promoting the degradation of the HBV Core protein through a Caspase-3-dependent pathway. We first show that RNF5 expression is upregulated in HBV-infected cells and patient samples, suggesting a role in the host's antiviral response. Subsequently, we investigate the mechanism by which RNF5 mediates its antiviral effect, finding that RNF5 targets the Core protein for degradation independently of its E3 ubiquitin ligase activity. The degradation of Core protein is mediated through a Caspase-3-dependent mechanism rather than the proteasomal pathway. Interestingly, RNF5's antiviral function does not rely on ubiquitination, indicating an alternative pathway involving apoptosis-related processes. These findings highlight the multifunctional role of RNF5 and suggest that targeting RNF5 could serve as a novel therapeutic approach to control HBV replication, providing new insights into the development of antiviral therapies against HBV.
Keywords: Caspase-3; E3 ubiqitin ligase; HBV - hepatitis B virus; RNF5; core protein.
Copyright © 2025 Xu, Song, Xu, Gao, Jiang and Tan.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
The E3 ligase RNF5 restricts SARS-CoV-2 replication by targeting its envelope protein for degradation.Signal Transduct Target Ther. 2023 Feb 3;8(1):53. doi: 10.1038/s41392-023-01335-5. Signal Transduct Target Ther. 2023. PMID: 36737599 Free PMC article.
-
Ring finger protein 5 mediates STING degradation through ubiquitinating K135 and K155 in a teleost fish.Front Immunol. 2024 Dec 11;15:1525376. doi: 10.3389/fimmu.2024.1525376. eCollection 2024. Front Immunol. 2024. PMID: 39723209 Free PMC article.
-
Swine RNF5 positively regulates the antiviral activity of IFITM1 by mediating the degradation of ABHD16A.J Virol. 2025 Jan 31;99(1):e0127724. doi: 10.1128/jvi.01277-24. Epub 2024 Nov 27. J Virol. 2025. PMID: 39601593 Free PMC article.
-
RNF5: inhibiting antiviral immunity and shaping virus life cycle.Front Immunol. 2024 Jan 5;14:1324516. doi: 10.3389/fimmu.2023.1324516. eCollection 2023. Front Immunol. 2024. PMID: 38250078 Free PMC article. Review.
-
Protein Degradation by Gammaherpesvirus RTAs: More Than Just Viral Transactivators.Viruses. 2023 Mar 11;15(3):730. doi: 10.3390/v15030730. Viruses. 2023. PMID: 36992439 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

