Tandem inactivation of inositol pyrophosphatases Asp1, Siw14, and Aps1 illuminates functional redundancies in inositol pyrophosphate catabolism in fission yeast
- PMID: 40237466
- PMCID: PMC12077094
- DOI: 10.1128/mbio.00389-25
Tandem inactivation of inositol pyrophosphatases Asp1, Siw14, and Aps1 illuminates functional redundancies in inositol pyrophosphate catabolism in fission yeast
Abstract
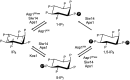
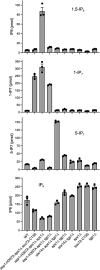
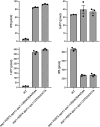
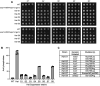
Inositol pyrophosphates 5-IP7, 1-IP7, and 1,5-IP8 are eukaryal signaling molecules that influence cell physiology, especially phosphate homeostasis. In fission yeast, 1,5-IP8 and 1-IP7 impact gene expression by acting as agonists of RNA 3'-processing and transcription termination. 1,5-IP8 is synthesized by position-specific kinases Kcs1 and Asp1 that convert IP6 to 5-IP7 and 5-IP7 to 1,5-IP8, respectively. Inositol pyrophosphatase enzymes Asp1 (a histidine acid phosphatase), Siw14 (a cysteinyl phosphatase), and Aps1 (a Nudix hydrolase) are agents of inositol pyrophosphate catabolism in fission yeast. Whereas Asp1, Siw14, and Aps1 are individually inessential, double pyrophosphatase mutants asp1-H397A aps1∆ and siw14∆ aps1∆ display severe growth defects caused by overzealous 3'-processing/termination. By applying CE-ESI-MS to profile the inositol pyrophosphate content of fission yeast mutants in which inositol pyrophosphate toxicity is genetically suppressed, we elucidated the functional redundancies of the Asp1, Siw14, and Aps1 pyrophosphatases. Asp1, which exclusively cleaves the 1-β-phosphate, and Aps1, which prefers to cleave the 1-β-phosphate, play essential overlapping roles in guarding against the accumulation of toxic levels of 1-IP7. Aps1 and Siw14 together catabolize the inositol-5-pyrophosphates, and their simultaneous inactivation results in overaccumulation of 5-IP7. Cells lacking all three pyrophosphatases amass high levels of 1,5-IP8 and 1-IP7, with concomitant depletion of IP6. A genetic screen identified three missense mutations in the catalytic domain of Kcs1 kinase that suppressed inositol-1-pyrophosphate toxicosis. The screen also implicated the 3'-processing factor Swd22, the inositol pyrophosphate sensor Spx1, and the nuclear poly(A)-binding protein Nab2 as mediators of inositol-1-pyrophosphate toxicity.IMPORTANCEInositol pyrophosphates are key effectors of eukaryal cellular phosphate homeostasis. They are synthesized by kinases that add a β-phosphate to the 5- or 1-phosphate groups of IP6 and catabolized by three classes of pyrophosphatases that hydrolyze the β-phosphates of 5-IP7, 1-IP7, or 1,5-IP8. Whereas the fission yeast inositol pyrophosphatases-Asp1 (histidine acid phosphatase), Siw14 (cysteinyl phosphatase), and Aps1 (Nudix hydrolase)-are inessential for growth, Asp1/Aps1 and Aps1/Siw14 double mutations and Asp1/Siw14/Aps1 triple mutations elicit severe or lethal growth defects. By profiling the inositol pyrophosphate content of pyrophosphatase mutants in which this toxicity is genetically suppressed, we reveal the functional redundancies of the Asp1, Siw14, and Aps1 pyrophosphatases. Their synergies are manifested as excess accumulation of 1-IP7 upon dual inactivation of Asp1 and Aps1 or an excess of 5-IP7 in aps1∆ siw14∆ cells. In the absence of all three pyrophosphatases, cells accrue high levels of 1,5-IP8 and 1-IP7 while IP6 declines.
Keywords: Schizosaccharomyces pombe; inositol pyrophosphates; pyrophosphatases.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Activities and genetic interactions of fission yeast Aps1, a Nudix-type inositol pyrophosphatase and inorganic polyphosphatase.mBio. 2024 Aug 14;15(8):e0108424. doi: 10.1128/mbio.01084-24. Epub 2024 Jun 28. mBio. 2024. PMID: 38940614 Free PMC article.
-
Genetic suppressor screen identifies Tgp1 (glycerophosphocholine transporter), Kcs1 (IP6 kinase), and Plc1 (phospholipase C) as determinants of inositol pyrophosphate toxicosis in fission yeast.mBio. 2024 Feb 14;15(2):e0306223. doi: 10.1128/mbio.03062-23. Epub 2023 Dec 22. mBio. 2024. PMID: 38133430 Free PMC article.
-
Activities, substrate specificity, and genetic interactions of fission yeast Siw14, a cysteinyl-phosphatase-type inositol pyrophosphatase.mBio. 2023 Oct 31;14(5):e0205623. doi: 10.1128/mbio.02056-23. Epub 2023 Sep 29. mBio. 2023. PMID: 37772819 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
References
-
- Pascual-Ortiz M, Saiardi A, Walla E, Jakopec V, Künzel NA, Span I, Vangala A, Fleig U. 2018. Asp1 bifunctional activity modulates spindle function via controlling cellular inositol pyrophosphate levels in Schizosaccharomyces pombe. Mol Cell Biol 38:e00047-18. doi:10.1128/MCB.00047-18 - DOI - PMC - PubMed
-
- Benjamin B, Garg A, Jork N, Jessen HJ, Schwer B, Shuman S. 2022. Activities and structure-function analysis of fission yeast inositol pyrophosphate (IPP) kinase-pyrophosphatase Asp1 and its impact of regulation of pho1 gene expression. mBio 13:e01034-22. doi:10.1128/mbio.01034-22 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

