Coupling of ribosome biogenesis and translation initiation in human mitochondria
- PMID: 40240327
- PMCID: PMC12003892
- DOI: 10.1038/s41467-025-58827-x
Coupling of ribosome biogenesis and translation initiation in human mitochondria
Abstract
Biogenesis of mitoribosomes requires dedicated chaperones, RNA-modifying enzymes, and GTPases, and defects in mitoribosome assembly lead to severe mitochondriopathies in humans. Here, we characterize late-step assembly states of the small mitoribosomal subunit (mtSSU) by combining genetic perturbation and mutagenesis analysis with biochemical and structural approaches. Isolation of native mtSSU biogenesis intermediates via a FLAG-tagged variant of the GTPase MTG3 reveals three distinct assembly states, which show how factors cooperate to mature the 12S rRNA. In addition, we observe four distinct primed initiation mtSSU states with an incompletely matured rRNA, suggesting that biogenesis and translation initiation are not mutually exclusive processes but can occur simultaneously. Together, these results provide insights into mtSSU biogenesis and suggest a functional coupling between ribosome biogenesis and translation initiation in human mitochondria.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures





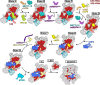
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

