Structure of the ATP-driven methyl-coenzyme M reductase activation complex
- PMID: 40240609
- PMCID: PMC12176620
- DOI: 10.1038/s41586-025-08890-7
Structure of the ATP-driven methyl-coenzyme M reductase activation complex
Abstract
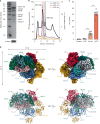
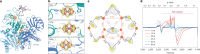
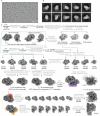
Methyl-coenzyme M reductase (MCR) is the enzyme responsible for nearly all biologically generated methane1. Its active site comprises coenzyme F430, a porphyrin-based cofactor with a central nickel ion that is active exclusively in the Ni(I) state2,3. How methanogenic archaea perform the reductive activation of F430 represents a major gap in our understanding of one of the most ancient bioenergetic systems in nature. Here we purified and characterized the MCR activation complex from Methanococcus maripaludis. McrC, a small subunit encoded in the mcr operon, co-purifies with the methanogenic marker proteins Mmp7, Mmp17, Mmp3 and the A2 component. We demonstrated that this complex can activate MCR in vitro in a strictly ATP-dependent manner, enabling the formation of methane. In addition, we determined the cryo-electron microscopy structure of the MCR activation complex exhibiting different functional states with local resolutions reaching 1.8-2.1 Å. Our data revealed three complex iron-sulfur clusters that formed an electron transfer pathway towards F430. Topology and electron paramagnetic resonance spectroscopy analyses indicate that these clusters are similar to the [8Fe-9S-C] cluster, a maturation intermediate of the catalytic cofactor in nitrogenase. Altogether, our findings offer insights into the activation mechanism of MCR and prospects on the early evolution of nitrogenase.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures










References
-
- Jackson, R. B. et al. Human activities now fuel two-thirds of global methane emissions. Environ. Res. Lett.19, 101002 (2024). - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

