Exploring methylation signatures for high de novo recurrence risk in hepatocellular carcinoma
- PMID: 40241383
- PMCID: PMC12016632
- DOI: 10.3350/cmh.2024.0899
Exploring methylation signatures for high de novo recurrence risk in hepatocellular carcinoma
Abstract
Background/aims: Hepatocellular carcinoma (HCC) exhibits high de novo recurrence rates post-resection. Current post-surgery recurrence prediction methods are limited, emphasizing the need for reliable biomarkers to assess recurrence risk. We aimed to develop methylation-based markers for classifying HCC patients and predicting their risk of de novo recurrence post-surgery.
Methods: In this retrospective cohort study, we analyzed data from HCC patients who underwent surgical resection in Korea, excluding those with recurrence within one year post-surgery. Using the Infinium Methylation EPIC array on 140 samples in the discovery cohort, we classified patients into low- and high-risk groups based on methylation profiles. Distinctive markers were identified through random forest analysis. These markers were validated in the cancer genome atlas (n=217), Validation cohort 1 (n=63) and experimental Validation using a methylation-sensitive high-resolution melting (MS-HRM) assay in Validation cohort 1 and Validation cohort 2 (n=63).
Results: The low-risk recurrence group (methylation group 1; MG1) showed a methylation average of 0.73 (95% confidence interval [CI] 0.69-0.77) with a 23.5% recurrence rate, while the high-risk group (MG2) had an average of 0.17 (95% CI 0.14-0.20) with a 44.1% recurrence rate (P<0.03). Validation confirmed the applicability of methylation markers across diverse populations, showing high accuracy in predicting the probability of HCC recurrence risk (area under the curve 96.8%). The MS-HRM assay confirmed its effectiveness in predicting de novo recurrence with 95.5% sensitivity, 89.7% specificity, and 92.2% accuracy.
Conclusion: Methylation markers effectively classified HCC patients by de novo recurrence risk, enhancing prediction accuracy and potentially offering personalized management strategies.
Keywords: Biomarker; DNA methylation; Hepatocellular carcinoma; Machine learning; Recurrence.
Conflict of interest statement
The authors have no conflicts to disclose.
Figures
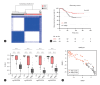



References
-
- Shibata T, Aburatani H. Exploration of liver cancer genomes. Nat Rev Gastroenterol Hepatol. 2014;11:340–349. - PubMed
-
- Mazzaferro V, Bhoori S, Sposito C, Bongini M, Langer M, Miceli R, et al. Milan criteria in liver transplantation for hepatocellular carcinoma: an evidence-based analysis of 15 years of experience. Liver Transpl. 2011;17 Suppl 2:S44–S57. - PubMed
-
- Lee SM, Kim-Ha J, Choi WY, Lee J, Kim D, Lee J, et al. Interplay of genetic and epigenetic alterations in hepatocellular carcinoma. Epigenomics. 2016;8:993–1005. - PubMed
-
- Sapisochin G, Bruix J. Liver transplantation for hepatocellular carcinoma: outcomes and novel surgical approaches. Nat Rev Gastroenterol Hepatol. 2017;14:203–217. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

