Circ_RUSC2 Sequesters miR-661 and Elevates TUSC2 Expression to Suppress Colorectal Cancer Progression
- PMID: 40243558
- PMCID: PMC11989122
- DOI: 10.3390/ijms26072937
Circ_RUSC2 Sequesters miR-661 and Elevates TUSC2 Expression to Suppress Colorectal Cancer Progression
Abstract
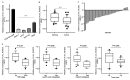
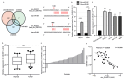
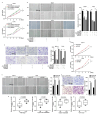
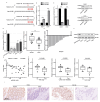
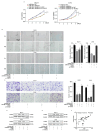
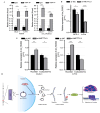
Despite advancements in diagnostic efficiency, colorectal cancer (CRC) remains a leading cause of cancer-related mortality, with increasing incidence rates. Circular RNA (circRNA) is a closed-loop, generally stable noncoding RNA that functions as a sponge for microRNAs in CRC. The purpose of this study was to investigate the function and underlying mechanism of circ_RUSC2, a new circRNA, in CRC. The expression levels of circ_RUSC2, miR-661, and TUSC2 were assessed using qRT-PCR, Western blot, and immunohistochemistry. Functional assays, including CCK-8, Transwell, and scratch wound healing, were performed to evaluate cell proliferation, migration, and invasion. RNA pull-down and actinomycin D assays were used to study RNA interactions and stability. In both CRC cells and tissues, miR-661 was markedly elevated, while circ_RUSC2 expression was considerably reduced. Poor differentiation, distant metastases, lymph node metastases, and an advanced stage were all strongly correlated with either miR-661 overexpression or circ_RUSC2 downregulation. circ_RUSC2 was more stable compared to its linear RUSC2 mRNA. CRC cell invasion, migration, and proliferation were suppressed by circ_RUSC2 ectopic expression; this inhibitory effect was restored by a miR-661 mimic. Circ_RUSC2 served as miR-661's sponge. TUSC2 counteracted the effects of miR-661, which stimulated CRC cell proliferation, migration, and invasion. At the post-transcriptional level, miR-661 controlled the expression of TUSC2 in CRC cells. In comparison to the negative control, circ_RUSC2 expression was markedly reduced, and its half-life was shortened by methyltransferase-like 3 (METTL3) knockdown. Circ_RUSC2 is a stable cytoplasmic circRNA. Circ_RUSC2 inhibits CRC cell malignant phenotypes via the miR-661/TUSC2 axis. The onset and progression of CRC are linked to the downregulation of Circ_RUSC2. circ_RUSC2 might become more stable through N6-methyladenosine (m6A) methylation regulated by METTL3. According to our research, circ_RUSC2 might be a new biomarker and treatment target for CRC.
Keywords: METTL3; TUSC2; circ_RUSC2; colorectal cancer; microRNA-661.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

Similar articles
-
Circ_0084615 is an oncogenic circular RNA in colorectal cancer and promotes DNMT3A expression via repressing miR-599.Pathol Res Pract. 2021 Aug;224:153494. doi: 10.1016/j.prp.2021.153494. Epub 2021 May 27. Pathol Res Pract. 2021. PMID: 34091391
-
EIF4A3-induced circ_0084615 contributes to the progression of colorectal cancer via miR-599/ONECUT2 pathway.J Exp Clin Cancer Res. 2021 Jul 12;40(1):227. doi: 10.1186/s13046-021-02029-y. J Exp Clin Cancer Res. 2021. PMID: 34253241 Free PMC article.
-
Circ_0000317/microRNA-520g/HOXD10 axis affects the biological characteristics of colorectal cancer.Kaohsiung J Med Sci. 2021 Nov;37(11):951-963. doi: 10.1002/kjm2.12422. Epub 2021 Jul 22. Kaohsiung J Med Sci. 2021. PMID: 34292663 Free PMC article.
-
Circular RNA (circ)_0053277 Contributes to Colorectal Cancer Cell Growth, Angiogenesis, Metastasis and Glycolysis.Mol Biotechnol. 2024 Nov;66(11):3285-3299. doi: 10.1007/s12033-023-00936-3. Epub 2023 Nov 2. Mol Biotechnol. 2024. PMID: 37917325
-
Circular RNA hsa_circ_0005939 Regulates UHRF1BP1L Expression by Targeting miR-4693-3p to Promote Colorectal Cancer Progression.Protein Pept Lett. 2024;31(6):437-446. doi: 10.2174/0109298665297110240611115010. Protein Pept Lett. 2024. PMID: 38918974
Cited by
-
Circular RNAs as Targets for Developing Anticancer Therapeutics.Cells. 2025 Jul 18;14(14):1106. doi: 10.3390/cells14141106. Cells. 2025. PMID: 40710359 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

