Active control of mitochondrial network morphology by metabolism-driven redox state
- PMID: 40244668
- PMCID: PMC12037031
- DOI: 10.1073/pnas.2421953122
Active control of mitochondrial network morphology by metabolism-driven redox state
Abstract
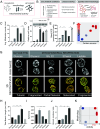
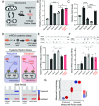
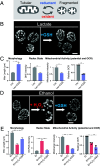
Mitochondria are dynamic organelles that constantly change morphology. What controls mitochondrial morphology however remains unresolved. Using actively respiring yeast cells growing in distinct carbon sources, we find that mitochondrial morphology and activity are unrelated. Cells can exhibit fragmented or networked mitochondrial morphology in different nutrient environments independent of mitochondrial activity. Instead, mitochondrial morphology is controlled by the intracellular redox state, which itself depends on the nature of electron entry into the electron transport chain (ETC)-through complex I/II or directly to coenzyme Q/cytochrome c. In metabolic conditions where direct electron entry is high, reactive oxygen species (ROS) increase, resulting in an oxidized cytosolic environment and rapid mitochondrial fragmentation. Decreasing direct electron entry into the ETC by genetic or chemical means, or reducing the cytosolic environment rapidly restores networked morphologies. Using controlled disruptions of electron flow to alter ROS and redox state, we demonstrate minute-scale, reversible control between networked and fragmented forms in an activity-independent manner. Mechanistically, the fission machinery through Dnm1 responds in minute-scale to redox state changes, preceding the change in mitochondrial form. Thus, the metabolic state of the cell and its consequent cellular redox state actively control mitochondrial form.
Keywords: electron transport chain; mitochondrial network; reactive oxygen species; redox state.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures



References
-
- Pelley J. W., “7—Citric acid cycle, electron transport chain, and oxidative phosphorylation” in Elsevier’s Integrated Review Biochemistry, Pelley J. W., Ed. (W.B. Saunders, ed. 2, 2012), pp. 57–65.
MeSH terms
Substances
Grants and funding
- DBT RA PDF/Department of Biotechnology, Ministry of Science and Technology, India (DBT)
- IA/S/21/2/505922/Wellcome Trust DBT India Alliance (India Alliance)
- IA/S/21/2/505922/WTDBT_/DBT-Wellcome Trust India Alliance/India
- WT_/Wellcome Trust/United Kingdom
- S. Ramachandran National Bioscience Award for Career Development/Department of Biotechnology, Ministry of Science and Technology, India (DBT)
LinkOut - more resources
Full Text Sources

