The Saccharomyces cerevisiae amino acid transporter Lyp1 has a broad substrate spectrum
- PMID: 40247776
- PMCID: PMC12147432
- DOI: 10.1002/1873-3468.70044
The Saccharomyces cerevisiae amino acid transporter Lyp1 has a broad substrate spectrum
Abstract
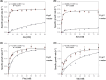
The main mediators for the amino acid uptake in Saccharomyces cerevisiae are the permeases belonging to the yeast amino acid transporter family. Recently, we discovered that members of this family support growth on more amino acids than previously described. Here we study the substrate spectrum of Lyp1, the main transporter responsible for the uptake of lysine in yeast. We show that overexpressed Lyp1 supports growth on alanine, asparagine, leucine, methionine, phenylalanine, serine, and valine when these are provided as the sole source of nitrogen to a strain severely deficient for the uptake of amino acids. We show that alanine and serine compete with lysine for the common transport system, albeit with much lower affinity. Thus, Lyp1 has a much broader substrate spectrum than previously thought, which may be true for many amino acid transporters.
Keywords: Saccharomyces cerevisiae; competitive inhibition; growth assay; low‐affinity substrate; lysine permease; yeast amino acid transporter.
© 2025 The Author(s). FEBS Letters published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
The authors disclose no conflict of interest.
Figures




Similar articles
-
Functional mapping and implications of substrate specificity of the yeast high-affinity leucine permease Bap2.Biochim Biophys Acta. 2014 Jul;1838(7):1719-29. doi: 10.1016/j.bbamem.2014.03.018. Epub 2014 Mar 31. Biochim Biophys Acta. 2014. PMID: 24699373
-
Amino acid residues important for substrate specificity of the amino acid permeases Can1p and Gnp1p in Saccharomyces cerevisiae.Yeast. 2001 Nov;18(15):1429-40. doi: 10.1002/yea.792. Yeast. 2001. PMID: 11746604
-
Extracellular loops matter - subcellular location and function of the lysine transporter Lyp1 from Saccharomyces cerevisiae.FEBS J. 2020 Oct;287(20):4401-4414. doi: 10.1111/febs.15262. Epub 2020 Mar 11. FEBS J. 2020. PMID: 32096906 Free PMC article.
-
Function and Regulation of Fungal Amino Acid Transporters: Insights from Predicted Structure.Adv Exp Med Biol. 2016;892:69-106. doi: 10.1007/978-3-319-25304-6_4. Adv Exp Med Biol. 2016. PMID: 26721271 Review.
-
Conservation of amino acid transporters in fungi, plants and animals.Trends Biochem Sci. 2002 Mar;27(3):139-47. doi: 10.1016/s0968-0004(01)02054-0. Trends Biochem Sci. 2002. PMID: 11893511 Review.
References
-
- Bender DA (2012) Amino Acid Metabolism. John Wiley & Sons, Incorporated.
-
- Barnett JA (2008) A history of research on yeasts 13. Active transport and the uptake of various metabolites. Yeast 25, 689–731. - PubMed
-
- Jack DL, Paulsen IT and Saier J (2000) The amino acid/polyamine/organocation (APC) superfamily of transporters specific for amino acids, polyamines and organocations. Microbiology 146, 1797–1814. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

