The complete chloroplast genome and phylogenetic analysis of Cinnamomum daphnoides (Laurales: Lauraceae)
- PMID: 40248043
- PMCID: PMC12004720
- DOI: 10.1080/23802359.2025.2492098
The complete chloroplast genome and phylogenetic analysis of Cinnamomum daphnoides (Laurales: Lauraceae)
Abstract
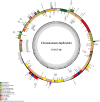
This study presents the complete chloroplast genome of Cinnamomum daphnoides (154,121 bp, GC 39.2%), a newly recorded species in China, revealing a typical quadripartite structure: a large single-copy region (93,688 bp, GC 37.9%), a small single-copy region (18,929 bp, GC 33.9%), and two inverted repeats (20,752 bp each, GC 44.3%). The genome harbors 126 genes (82 protein-coding, 36 tRNA, 8 rRNA). Phylogenetic analysis groups C. daphnoides with C. tamala, differing from prior studies. These findings clarify its systematic position within Cinnamomum and support its potential for coastal greening applications.
Keywords: Cinnamomum daphnoides; Lauraceae; chloroplast genome; phylogeny.
© 2025 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group.
Conflict of interest statement
No potential conflict of interest was reported by the authors.
Figures


References
-
- Jinlu L, Shuo W, Jing Y, Ling W, Shiliang Z.. 2013. A modified CTAB protocol for plant DNA extraction. Chinese Bulletin Of Botany. 48(1):72–78. doi:10.3724/SP.J.1259.2013.00072. - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
