Integrated liver-secreted and plasma proteomics identify a predictive model that stratifies MASH
- PMID: 40250425
- PMCID: PMC12147855
- DOI: 10.1016/j.xcrm.2025.102085
Integrated liver-secreted and plasma proteomics identify a predictive model that stratifies MASH
Abstract
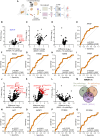
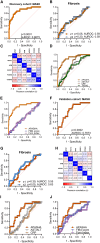
Obesity is a major risk factor for metabolic-associated steatotic liver disease (MASLD), which can progress to metabolic-associated steatohepatitis (MASH). There are no validated non-invasive tests to stratify persons with obesity with a greater risk for MASH. Herein, we assess plasma and liver from 266 obese individuals spanning the MASLD spectrum. Ninety-six human livers were precision-cut, and mass spectrometry-based proteomics identifies 3,333 proteins in the liver-secretion medium, of which 107 are differentially secreted in MASH compared with no pathology. The plasma proteome is markedly remodeled in MASH but is not different between patients with steatosis and no pathology. The APASHA model, comprising plasma apolipoprotein F (APOF), proprotein convertase subtilisin/kexin type 9 (PCSK9), afamin (AFM), S100 calcium-binding protein A6 (S100A6), HbA1c, and zinc-alpha-2-glycoprotein (AZGP1), stratifies MASH (area under receiver operating characteristic [AUROC] = 0.88). Our investigations detail the evolution of liver-secreted and plasma proteins with MASLD progression, providing a rich resource defining human liver-secreted proteins and creating a predictive model to stratify patients with obesity at risk of MASH.
Keywords: APASHA; APOF; AZGP1; MASL; MASLD; hepatokine; protein secretion; proteome; small protein enrichment.
Copyright © 2025 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests W.A.B. reports financial support for a bariatric surgery registry from the Commonwealth of Australia, Apollo Endosurgery, Covidien, Johnson and Johnson, Gore, and Applied Medical. She has also received a speaker’s honorarium from Merck Sharp and Dohme and a speaker’s honorarium and fees for participation in a scientific advisory board from Novo Nordisk. The Bariatric Registry and the honorariums are outside of the submitted work. M.J.W. has received financial support from Gilead Sciences and CSL.
Figures



Similar articles
-
Liver Tissue Proteins Improve the Accuracy of Plasma Proteins as Biomarkers in Diagnosing Metabolic Dysfunction-Associated Steatohepatitis.Proteomics Clin Appl. 2024 Nov;18(6):e202300236. doi: 10.1002/prca.202300236. Epub 2024 Jul 28. Proteomics Clin Appl. 2024. PMID: 39073724
-
One-hour post-load glucose is associated with biopsy-proven metabolic dysfunction-associated steatotic liver disease in obese individuals.Diabetes Metab. 2025 Jan;51(1):101588. doi: 10.1016/j.diabet.2024.101588. Epub 2024 Dec 5. Diabetes Metab. 2025. PMID: 39645036
-
Serum lipidomic signatures in patients with varying histological severity of metabolic-dysfunction associated steatotic liver disease.Metabolism. 2025 Jan;162:156063. doi: 10.1016/j.metabol.2024.156063. Epub 2024 Nov 8. Metabolism. 2025. PMID: 39522592
-
Uncovering hepatic transcriptomic and circulating proteomic signatures in MASH: A meta-analysis and machine learning-based biomarker discovery.Comput Biol Med. 2025 Jun;191:110170. doi: 10.1016/j.compbiomed.2025.110170. Epub 2025 Apr 12. Comput Biol Med. 2025. PMID: 40220593
-
NAFLD (MASLD)/NASH (MASH): Does It Bother to Label at All? A Comprehensive Narrative Review.Int J Mol Sci. 2024 Aug 2;25(15):8462. doi: 10.3390/ijms25158462. Int J Mol Sci. 2024. PMID: 39126031 Free PMC article. Review.
References
-
- Estes C., Anstee Q.M., Arias-Loste M.T., Bantel H., Bellentani S., Caballeria J., Colombo M., Craxi A., Crespo J., Day C.P., et al. Modeling NAFLD disease burden in China, France, Germany, Italy, Japan, Spain, United Kingdom, and United States for the period 2016–2030. J. Hepatol. 2018;69:896–904. doi: 10.1016/j.jhep.2018.05.036. - DOI - PubMed
-
- Kleiner D.E., Brunt E.M., Van Natta M., Behling C., Contos M.J., Cummings O.W., Ferrell L.D., Liu Y.C., Torbenson M.S., Unalp-Arida A., et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology. 2005;41:1313–1321. doi: 10.1002/hep.20701. - DOI - PubMed
-
- Singh S., Allen A.M., Wang Z., Prokop L.J., Murad M.H., Loomba R. Fibrosis progression in nonalcoholic fatty liver vs nonalcoholic steatohepatitis: a systematic review and meta-analysis of paired-biopsy studies. Clin. Gastroenterol. Hepatol. 2015;13:643–654.e1-9. doi: 10.1016/j.cgh.2014.04.014. quiz e39-40. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

