Transcription factor JUNB is required for transformation of EpCAM-positive hepatocellular carcinoma (HCC) cells into CD90-positive HCC cells in vitro
- PMID: 40253402
- PMCID: PMC12009367
- DOI: 10.1038/s41419-025-07602-3
Transcription factor JUNB is required for transformation of EpCAM-positive hepatocellular carcinoma (HCC) cells into CD90-positive HCC cells in vitro
Abstract
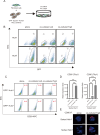
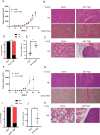
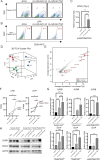
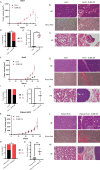
Hepatocellular carcinoma (HCC) harbors two types of stem cells-epithelial and mesenchymal stem cells. The mechanism by which epithelial EpCAM-positive HCC cells transform into mesenchymal CD90-positive HCC cells remains unclear. On peritumoral fibrotic nodules, epithelial HCC cells form communities with stromal cells, driving tumor growth and malignancy. We aimed to clarify the mechanism by which epithelial cell adhesion molecule (EpCAM)-positive HCC cells contribute to the phenotype of mesenchymal CD90-positive HCC cells that metastasize to distant sites by elucidating the interaction between EpCAM-positive HCC cells and fibroblasts. EpCAM-positive CD90-negative epithelial HCC cells (Huh1, Huh7, and HCC cells) were converted into metastasis-prone CD90-positive HCC cells by co-culture with fibroblasts (Lx-2 and Tig3-20). We identified the transcription factor JUNB as responsible for this altered phenotype. We found that the overexpression of JUNB in CD90-negative epithelial HCC cells resulted in significant transformation to mesenchymal CD90-positive HCC in vitro and in vivo, showing metastatic potential to the lungs. In addition, the JUNB expression in EpCAM-positive hepatoma cells was increased by paracrine stimulation with fibroblast-derived TGFb1. This study unravels the mechanism by which fibroblasts aggravate the malignancy of liver cancer, and the results suggest that JUNB may be a target for treating liver cancer metastasis.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. Ethics approval and consent to participate: All animal experiments were approved by the Animal Care and Use Committees of the Institute of Medical, Pharmaceutical, and Health Sciences, Kanazawa University. They were also conducted following the institutional ethical guidelines for animal experiments and the safety guidelines for gene manipulation experiments. The clinical specimens used in this study were performed in accordance with the Declaration of Helsinki, and the protocol was approved by the Ethics Committee of Kanazawa University Graduate School of Medicine. All subjects gave their informed consent for inclusion before they participated in the study, in which the tissue would be used for future study.
Figures

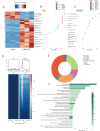


Similar articles
-
Sorafenib suppresses extrahepatic metastasis de novo in hepatocellular carcinoma through inhibition of mesenchymal cancer stem cells characterized by the expression of CD90.Sci Rep. 2017 Sep 12;7(1):11292. doi: 10.1038/s41598-017-11848-z. Sci Rep. 2017. PMID: 28900199 Free PMC article.
-
Discrete nature of EpCAM+ and CD90+ cancer stem cells in human hepatocellular carcinoma.Hepatology. 2013 Apr;57(4):1484-97. doi: 10.1002/hep.26168. Epub 2013 Jan 18. Hepatology. 2013. PMID: 23174907 Free PMC article.
-
Expression of Cancer Stem Cell Markers EpCAM and CD90 Is Correlated with Anti- and Pro-Oncogenic EphA2 Signaling in Hepatocellular Carcinoma.Int J Mol Sci. 2021 Aug 11;22(16):8652. doi: 10.3390/ijms22168652. Int J Mol Sci. 2021. PMID: 34445353 Free PMC article.
-
Cellular and Molecular Biology of Cancer Stem Cells of Hepatocellular Carcinoma.Int J Mol Sci. 2023 Jan 11;24(2):1417. doi: 10.3390/ijms24021417. Int J Mol Sci. 2023. PMID: 36674932 Free PMC article. Review.
-
Epithelial Mesenchymal and Endothelial Mesenchymal Transitions in Hepatocellular Carcinoma: A Review.Biomed Res Int. 2019 Sep 29;2019:2962580. doi: 10.1155/2019/2962580. eCollection 2019. Biomed Res Int. 2019. PMID: 31781608 Free PMC article. Review.
References
-
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49. 10.3322/caac.21660. - PubMed
-
- Villanueva A. Hepatocellular carcinoma. N Engl J Med. 2019;380:1450–62. 10.1056/NEJMra1713263. - PubMed
-
- Tabrizian P, Jibara G, Shrager B, Schwartz M, Roayaie S. Recurrence of hepatocellular cancer after resection: patterns, treatments, and prognosis. Ann Surg. 2015;261:947–55. 10.1097/SLA.0000000000000710. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

