RNA editing generates mRNA isoforms with distinct stabilities that may expand the thermal tolerance of mRNA and proteins in Mytilus species
- PMID: 40259733
- PMCID: PMC12361899
- DOI: 10.24272/j.issn.2095-8137.2024.383
RNA editing generates mRNA isoforms with distinct stabilities that may expand the thermal tolerance of mRNA and proteins in Mytilus species
Abstract
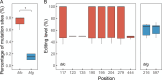
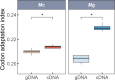
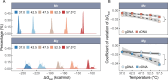
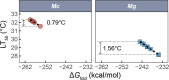
Ectothermic organisms may expand their thermal tolerance by producing multiple protein isoforms with differing thermal sensitivities. While such isoforms commonly originate from allelic variation at a single locus (allozymes) or from gene duplication that gives rise to paralogs with distinct thermal responses, this study investigated mRNA editing as an alternative, post-transcriptional mechanism for generating mRNA variants. Cytosolic malate dehydrogenase (cMDH) was examined in foot tissue of two congeners of the marine mussel genus Mytilus, which occupy different thermal environments. Multiple editing events were detected within the mRNA coding region in both species. Editing sites were species-specific, with no shared positions identified. In M. coruscus, editing occurred at 117, 123, 135, 190, 195, 204, 279, and 444, while in M. galloprovincialis, editing was detected at 216 and 597. Each species exhibited multiple edited mRNA variants, and these isoforms were associated with differential protein expression. These findings suggest that mRNA editing may contribute an additional layer of molecular variation. The generation of diverse mRNA isoforms from a single DNA coding sequence may enhance enzymatic flexibility across temperature ranges, supporting eurythermal physiological performance and mitigating thermal stress. Moreover, the presence of multiple edited transcripts within individual organisms raises important caveats about the limitations of approaches that deduce amino acid sequences or estimate adaptive variation solely from genomic data.
变温动物体内的蛋白质通过产生一系列热敏感性迥异的变体(同源异构体),扩大了机体的热耐受范围。从遗传学上讲,在响应高温过程中功能和结构稳定性上存在差异的同源异构体,起源于等位基因变异与基因复制。该研究以两种海洋贝类( Mytilus属)为研究对象,聚焦其产生mRNA变异的转录后调控机制——mRNA编辑的作用,探索了细胞质苹果酸脱氢酶(cMDH)的温度响应策略。研究结果表明,两种贻贝cMDH mRNA的编码序列区域均发生了RNA编辑事件。RNA编辑位点具有物种特异性,两个物种间不存在相同的编辑位点。对于厚壳贻贝( M. coruscus)而言,cMDH的 #117、#123、#135、#190、#195、#204、#279和#444位点发生了RNA编辑,而紫贻贝( M. galloprovincialis)cMDH的RNA编辑发生于#216和#597位点。经过RNA编辑产生了cMDH mRNA变体,分析结果表明上述mRNA变体在蛋白质表达上存在差异。基于此推断,由mRNA编辑引起的变异可以补充其他变异来源。这些由单一DNA编码位点变异所产生的mRNA同源异构体,可能促进了生物的生化功能向更广温的方向进化,增强了生物个体应对热胁迫的能力。此外,同一个体中的蛋白质可能存在由于RNA编辑导致的多种变体,在推导氨基酸序列和估计自然群体中的适应性分子变异时需重点关注。.
Keywords: RNA editing; Temperature adaptation; mRNA isoform; mRNA stability.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
-
- Bai ZY, Han XK, Luo M, et al Constructing a microsatellite-based linkage map and identifying QTL for pearl quality traits in triangle pearl mussel (Hyriopsis cumingii) Aquaculture. 2015;437:102–110. doi: 10.1016/j.aquaculture.2014.11.008. - DOI
-
- Bartoszewski RA, Jablonsky M, Bartoszewska S, et al A synonymous single nucleotide polymorphism in ΔF508 CFTR alters the secondary structure of the mRNA and the expression of the mutant protein. Journal of Biological Chemistry. 2010;285(37):28741–28748. doi: 10.1074/jbc.M110.154575. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
