Viral metagenomics reveals diverse viruses in the fecal samples of children with acute respiratory infection
- PMID: 40260089
- PMCID: PMC12009832
- DOI: 10.3389/fmicb.2025.1564755
Viral metagenomics reveals diverse viruses in the fecal samples of children with acute respiratory infection
Abstract
Introduction: Changes in the gut microbiome have been associated with the development of acute respiratory infection (ARI). However, due to methodological limitations, our knowledge of the gut virome in patients with ARIs remains limited.
Methods: In this study, fecal samples from children with ARI were investigated using viral metagenomics.
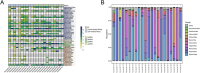
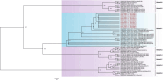
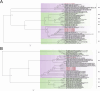
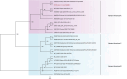
Results: The fecal virome was analyzed, and several suspected disease-causing viruses were identified. The five viral families with the highest abundance of sequence reads were Podoviridae, Virgaviridae, Siphoviridae, Microviridae, and Myoviridae. Additionally, human adenovirus, human bocavirus, human astrovirus, norovirus, and human rhinovirus were detected. The genome sequences of these viruses were respectively described, and phylogenetic trees were constructed using the gene sequences of the viruses.
Discussion: We characterized the composition of gut virome in children with acute respiratory infections. However, further research is required to elucidate the relationship between acute respiratory infection and gut viruses.
Keywords: acute respiratory infection; children; fecal samples; viral metagenomics; virus evolution.
Copyright © 2025 Xu, Pan, Yuan, Zhu, Wei, Lu and Zhang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Viral Metagenomic Analysis of the Fecal Samples in Domestic Dogs (Canis lupus familiaris).Viruses. 2023 Mar 6;15(3):685. doi: 10.3390/v15030685. Viruses. 2023. PMID: 36992396 Free PMC article.
-
Viral metagenomics analysis of feces from coronary heart disease patients reveals the genetic diversity of the Microviridae.Virol Sin. 2017 Apr;32(2):130-138. doi: 10.1007/s12250-016-3896-0. Epub 2017 Apr 17. Virol Sin. 2017. PMID: 28466442 Free PMC article.
-
Viral metagenomics of the gut virome of diarrheal children with Rotavirus A infection.Gut Microbes. 2023 Jan-Dec;15(1):2234653. doi: 10.1080/19490976.2023.2234653. Gut Microbes. 2023. PMID: 37448101 Free PMC article.
-
Insights Into the Role of the Lung Virome During Respiratory Viral Infections.Front Immunol. 2022 Apr 27;13:885341. doi: 10.3389/fimmu.2022.885341. eCollection 2022. Front Immunol. 2022. PMID: 35572506 Free PMC article. Review.
-
The Emerging Role of the Gut Virome in Health and Inflammatory Bowel Disease: Challenges, Covariates and a Viral Imbalance.Viruses. 2023 Jan 6;15(1):173. doi: 10.3390/v15010173. Viruses. 2023. PMID: 36680214 Free PMC article. Review.
References
-
- Alves J., Teixeira D., Siqueira J., Deus D., Oliveira D., Ferreira J., et al. (2024). Epidemiology and molecular detection of human adenovirus and non-polio enterovirus in fecal samples of children with acute gastroenteritis: A five-year surveillance in northern Brazil. PLoS One 19:e0296568. 10.1371/journal.pone.0296568 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

