Identification of a critical gene involved in the biosynthesis of the polyene macrolide lavencidin in Streptomyces lavendulae FRI-5 using the Target-AID (activation-induced cytidine deaminase) base editing technology
- PMID: 40261024
- PMCID: PMC12093942
- DOI: 10.1128/aem.00975-24
Identification of a critical gene involved in the biosynthesis of the polyene macrolide lavencidin in Streptomyces lavendulae FRI-5 using the Target-AID (activation-induced cytidine deaminase) base editing technology
Abstract
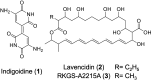
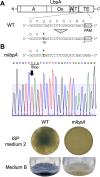
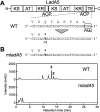
Polyene macrolide antibiotics, produced mainly as secondary metabolites of streptomycetes, have distinct chemical structures and include clinically important antifungal drugs. We recently isolated the 28-membered polyene macrolide lavencidin from Streptomyces lavendulae FRI-5. Here, we identify and characterize the lavencidin biosynthetic (lad) gene cluster by combining a gene disruption system based on a base editing technology and in silico analysis. Sequence analysis of the draft genome of S. lavendulae FRI-5 revealed plausible lavencidin biosynthetic genes, which could be assigned roles in the biosynthesis of the polyketide backbone and the peripheral moiety, as well as in the regulation of lavencidin production. The introduction of a stop codon into the ladA5 polyketide synthase (PKS) gene by the base editing system resulted in a complete loss of lavencidin production, indicating that the type I modular PKS system is responsible for the biosynthesis of lavencidin.IMPORTANCEPolyene macrolide antibiotics display a unique mode of action among fungicides and exhibit potent fungicidal activity to which resistance does not readily develop. Deciphering the biosynthetic pathways of these fascinating compounds will provide chemical diversity for the development of industrially and clinically important agents. In this study, the Target-AID (activation-induced cytidine deaminase) system enabled us to identify the lad gene cluster involved in lavencidin biosynthesis, paving the way for the rational design of lavencidin derivatives with new or improved biological activity. Furthermore, this base editing system is capable of precisely and rapidly substituting the target nucleotide in several streptomycetes. Thus, our Target-AID system would be a powerful and versatile tool for the genetic engineering of streptomycetes as well as for analyzing the functions of uncharacterized genes, expanding the chemical diversity of useful bioactive compounds, and discovering novel natural products.
Keywords: Streptomyces; base editing; lavencidin; polyene macrolide antibiotics; target-AID.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Zarins-Tutt JS, Barberi TT, Gao H, Mearns-Spragg A, Zhang L, Newman DJ, Goss RJM. 2016. Prospecting for new bacterial metabolites: a glossary of approaches for inducing, activating and upregulating the biosynthesis of bacterial cryptic or silent natural products. Nat Prod Rep 33:54–72. doi: 10.1039/c5np00111k - DOI - PubMed
-
- Hashimoto K, Nihira T, Sakuda S, Yamada Y. 1992. IM-2, a butyrolactone autoregulator, induces production of several nucleoside antibiotics in Streptomyces sp. FRI-5. J Ferment Bioeng 73:449–455. doi: 10.1016/0922-338X(92)90136-I - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

