Variant effect predictor correlation with functional assays is reflective of clinical classification performance
- PMID: 40264194
- PMCID: PMC12016141
- DOI: 10.1186/s13059-025-03575-w
Variant effect predictor correlation with functional assays is reflective of clinical classification performance
Abstract
Background: Understanding the relationship between protein sequence and function is crucial for accurate classification of missense variants. Variant effect predictors (VEPs) play a vital role in deciphering this complex relationship, yet evaluating their performance remains challenging for several reasons, including data circularity, where the same or related data is used for training and assessment. High-throughput experimental strategies like deep mutational scanning (DMS) offer a promising solution.
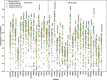
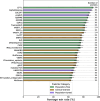
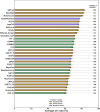
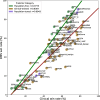
Results: In this study, we extend upon our previous benchmarking approach, assessing the performance of 97 VEPs using missense DMS measurements from 36 different human proteins. In addition, a new pairwise, VEP-centric approach mitigates the impact of missing predictions on overall performance comparison. We observe a strong correspondence between VEP performance in DMS-based benchmarks and clinical variant classification, especially for predictors that have not been directly trained on human clinical variants.
Conclusions: Our results suggest that comparing VEP performance against diverse functional assays represents a reliable strategy for assessing their relative performance in clinical variant classification. However, major challenges in clinical interpretation of VEP scores persist, highlighting the need for further research to fully leverage computational predictors for genetic diagnosis. We also address practical considerations for end users in terms of choice of methodology.
Keywords: ACMG/AMP; Benchmark; Circularity; DMS; MAVE; Multiplexed assay of variant effect; VEP; Variant effect predictor.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate.: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures
References
-
- Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–23. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

