Association Between Vascular NOTCH3 Aggregation and Disease Severity in a CADASIL Cohort - Implications for NOTCH3 Variant-Specific Disease Prediction
- PMID: 40265482
- PMCID: PMC12278194
- DOI: 10.1002/ana.27240
Association Between Vascular NOTCH3 Aggregation and Disease Severity in a CADASIL Cohort - Implications for NOTCH3 Variant-Specific Disease Prediction
Abstract
Objective: Vascular NOTCH3 protein ectodomain aggregation is a pathological hallmark of cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL), a monogenic small vessel disease typically caused by cysteine-altering variants in NOTCH3. Given their high population frequency, these NOTCH3 variants are an important genetic contributor to stroke and vascular dementia worldwide. Disease severity in CADASIL is highly variable and is mainly determined by the position of the pathogenic NOTCH3 variant in the NOTCH3 ectodomain. Here, we aimed to investigate the association between NOTCH3 aggregation load in skin vessels, cysteine-altering NOTCH3 variants, and disease severity in a prospective cohort study of 212 patients with CADASIL with 39 distinct cysteine-altering NOTCH3 variants.
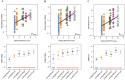
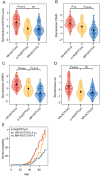
Methods: NOTCH3 aggregation load in skin vessels was determined by calculating the NOTCH3 score; the fraction of skin vessel wall area positive for NOTCH3 staining. Variant-specific NOTCH3 scores were calculated for variants present in 10 or more participants, by averaging the NOTCH3 scores of individuals with that distinct variant. The associations between the NOTCH3 score, NOTCH3 variants, and neuroimaging and clinical outcomes were investigated using multivariable linear mixed models, Cox regression, and mediation analyses.
Results: The NOTCH3 score was significantly associated with lifetime stroke probability and small vessel disease neuroimaging outcomes, but not with age. Variant-specific NOTCH3 scores reflected differences in disease severity associated with distinct NOTCH3 variants.
Interpretation: These findings suggest that differences in NOTCH3 aggregation propensity underlie the differences in disease severity associated with NOTCH3 cysteine-altering variants, and show that NOTCH3-variant specific NOTCH3 scores can contribute to improved individualized disease prediction in CADASIL. ANN NEUROL 2025;98:273-285.
© 2025 The Author(s). Annals of Neurology published by Wiley Periodicals LLC on behalf of American Neurological Association.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Joutel A, Vahedi K, Corpechot C, et al. Strong clustering and stereotyped nature of Notch3 mutations in CADASIL patients. Lancet 1997;350:1511–1515. - PubMed
-
- Rutten JW, Haan J, Terwindt GM, et al. Interpretation of NOTCH3 mutations in the diagnosis of CADASIL. Expert Rev Mol Diagn 2014;14:593–603. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

