Defining essential charged residues in fibril formation of a lysosomal derived N-terminal α-synuclein truncation
- PMID: 40268916
- PMCID: PMC12019160
- DOI: 10.1038/s41467-025-58899-9
Defining essential charged residues in fibril formation of a lysosomal derived N-terminal α-synuclein truncation
Abstract
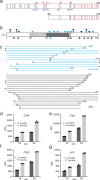
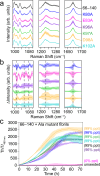
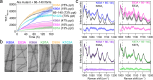
N- and C-terminal α-synuclein (α-syn) truncations are prevalent in Parkinson's disease. Effects of the N- and C-terminal residues on α-syn aggregation and fibril propagation are distinct, where the N-terminus dictates fibril structure. Here, the majority of α-syn truncations are assigned by intact mass spectrometry to lysosomal activity. To delineate essential charged residues in fibril formation, we selected an N-terminal truncation (66-140) that is generated solely from soluble α-syn by asparagine endopeptidase. Ala-substitutions at K80 and E83 impact aggregation kinetics, revealing their vital roles in defining fibril polymorphism. K80, E83, and K97 are identified to be critical for fibril elongation. Based on solid-state NMR, mutational and Raman studies, and molecular dynamics simulations, a E83-K97 salt bridge is proposed. Finally, participation of C-terminal Lys residues in the full-length α-syn fibril assembly process is also shown, highlighting that individual residues can be targeted for therapeutic intervention.
© 2025. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures





References
-
- Wakabayashi, K. et al. Accumulation of α-synuclein/NACP is a cytopathological feature common to Lewy body disease and multiple system atrophy. Acta Neuropathol.96, 445–452 (1998). - PubMed
-
- Scheres, S. H. W., Ryskeldi-Falcon, B. & Goedert, M. Molecular pathology of neurodegenerative diseases by cryo-EM of amyloids. Nature621, 701–710 (2023). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

