Regulation of pericentromeric DNA loop size via Scc2-cohesin interaction
- PMID: 40271018
- PMCID: PMC12017868
- DOI: 10.1016/j.isci.2025.112322
Regulation of pericentromeric DNA loop size via Scc2-cohesin interaction
Abstract
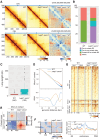
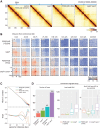
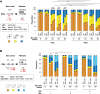
Cohesin exhibits DNA loop extrusion when bound to the ATPase activator Scc2 (NIPBL in humans), which has been proposed to organize higher-order chromosome folding. In budding yeast, most chromosome-bound cohesins lack Scc2. How the Scc2-cohesin interaction is regulated on the chromosome and its physiological consequences remain unclear. Here, we show that the deletion of both ECO1 and WPL1, two known cohesin regulators, but not either alone, caused Scc2-cohesin co-localization in metaphase, particularly around centromeres, using calibrated chromatin immunoprecipitation sequencing (ChIP-seq). Eco1's mitotic activity was required to prevent this co-localization in Δwpl1. We also demonstrate that Scc2-cohesin co-localization enlarged pericentromeric DNA loops, linking centromeres to genome sites hundreds of kilobases away, and delayed mitotic chromosome segregation. These findings suggest that Wpl1 and Eco1 cooperatively regulate Scc2-cohesin interaction, restrict pericentromeric DNA loop size, and facilitate chromosome segregation.
Keywords: Natural sciences; biological sciences; cell biology; functional aspects of cell biology.
© 2025 The Authors.
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Lieberman-Aiden E., Van Berkum N.L., Williams L., Imakaev M., Ragoczy T., Telling A., Amit I., Lajoie B.R., Sabo P.J., Dorschner M.O., et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–293. doi: 10.1126/SCIENCE.1181369. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

