Generation of a compound heterozygous ABCA4 rat model with pathological features of STGD1
- PMID: 40273359
- PMCID: PMC12138338
- DOI: 10.1093/hmg/ddaf057
Generation of a compound heterozygous ABCA4 rat model with pathological features of STGD1
Abstract
The ABCA4 protein plays an essential role in mammalian vision, ensuring the correct localization of all-trans-retinal within the visual cycle. Mutations in the ABCA4 gene are responsible for the juvenile maculopathy, Stargardt disease (STGD1). We investigated the most common variant underlying STGD1 phenotype in a rat model carrying the ortholog to the human c.5882G > A/p.(Gly1961Glu) (G1961E) in ABCA4. While the pathogenicity of this variant has recently been questioned, we examine here whether the ortholog rat variant is associated with vitamin A toxicity in the retina. By crossing the rat line with a rat line deficient in ABCA4 protein, we reveal a more pathogenic phenotype in line with compound heterozygosity, making the model suitable for testing of gene, cell and pharmacological therapies.
Keywords: ATPase Binding Cassette Transporter 4; G1961E variant; Stargardt disease; genetically modified rat.
© The Author(s) 2025. Published by Oxford University Press.
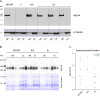
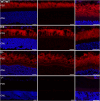
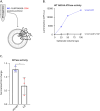
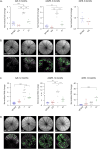
Figures




Similar articles
-
Identification of novel pathogenic ABCA4 variants in a Han Chinese family with Stargardt disease.Biosci Rep. 2019 Jan 15;39(1):BSR20180872. doi: 10.1042/BSR20180872. Print 2019 Jan 31. Biosci Rep. 2019. PMID: 30563929 Free PMC article.
-
Correlating the Expression and Functional Activity of ABCA4 Disease Variants With the Phenotype of Patients With Stargardt Disease.Invest Ophthalmol Vis Sci. 2018 May 1;59(6):2305-2315. doi: 10.1167/iovs.17-23364. Invest Ophthalmol Vis Sci. 2018. PMID: 29847635 Free PMC article.
-
Localization and functional characterization of the p.Asn965Ser (N965S) ABCA4 variant in mice reveal pathogenic mechanisms underlying Stargardt macular degeneration.Hum Mol Genet. 2018 Jan 15;27(2):295-306. doi: 10.1093/hmg/ddx400. Hum Mol Genet. 2018. PMID: 29145636 Free PMC article.
-
Proposing Zebrafish as a Model for Stargardt Disease.Adv Exp Med Biol. 2025;1468:219-223. doi: 10.1007/978-3-031-76550-6_36. Adv Exp Med Biol. 2025. PMID: 39930199 Review.
-
Gene Therapy of ABCA4-Associated Diseases.Cold Spring Harb Perspect Med. 2015 Jan 8;5(5):a017301. doi: 10.1101/cshperspect.a017301. Cold Spring Harb Perspect Med. 2015. PMID: 25573774 Free PMC article. Review.
References
-
- Higgins CF. ABC transporters: from microorganisms to man. Annu Rev Cell Biol 1992;8:67–113. - PubMed
-
- Sun H, Nathans J. Stargardt’s ABCR is localized to the disc membrane of retinal rod outer segments. Nat Genet 1997;17:15–16. - PubMed
-
- Molday RS, Garces FA, Scortecci JF. et al. Structure and function of ABCA4 and its role in the visual cycle and Stargardt macular degeneration. Prog Retin Eye Res 2022;89:101036. - PubMed
-
- Eldred GE, Lasky MR. Retinal age pigments generated by self-assembling lysosomotropic detergents. Nature 1993;361:724–726. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

