Endoplasmic Reticulum Stress and Its Role in Metabolic Reprogramming of Cancer
- PMID: 40278350
- PMCID: PMC12029571
- DOI: 10.3390/metabo15040221
Endoplasmic Reticulum Stress and Its Role in Metabolic Reprogramming of Cancer
Abstract
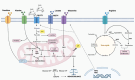
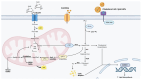
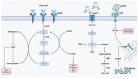
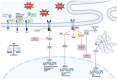
Background/Objectives: Endoplasmic reticulum (ER) stress occurs when ER homeostasis is disrupted, leading to the accumulation of misfolded or unfolded proteins. This condition activates the unfolded protein response (UPR), which aims to restore balance or trigger cell death if homeostasis cannot be achieved. In cancer, ER stress plays a key role due to the heightened metabolic demands of tumor cells. This review explores how metabolomics can provide insights into ER stress-related metabolic alterations and their implications for cancer therapy. Methods: A comprehensive literature review was conducted to analyze recent findings on ER stress, metabolomics, and cancer metabolism. Studies examining metabolic profiling of cancer cells under ER stress conditions were selected, with a focus on identifying potential biomarkers and therapeutic targets. Results: Metabolomic studies highlight significant shifts in lipid metabolism, protein synthesis, and oxidative stress management in response to ER stress. These metabolic alterations are crucial for tumor adaptation and survival. Additionally, targeting ER stress-related metabolic pathways has shown potential in preclinical models, suggesting new therapeutic strategies. Conclusions: Understanding the metabolic impact of ER stress in cancer provides valuable opportunities for drug development. Metabolomics-based approaches may help identify novel biomarkers and therapeutic targets, enhancing the effectiveness of antitumor therapies.
Keywords: biochemical pathways; cancer; drug discovery; endoplasmic reticulum stress; metabolomics; tumor microenvironment; unfolded protein response.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
The role of endoplasmic reticulum stress in promoting aerobic glycolysis in cancer cells: An overview.Pathol Res Pract. 2023 Nov;251:154905. doi: 10.1016/j.prp.2023.154905. Epub 2023 Oct 24. Pathol Res Pract. 2023. PMID: 37925820 Review.
-
Misfolded proteins bind and activate death receptor 5 to trigger apoptosis during unresolved endoplasmic reticulum stress.Elife. 2020 Jan 6;9:e52291. doi: 10.7554/eLife.52291. Elife. 2020. PMID: 31904339 Free PMC article.
-
Endoplasmic reticulum stress and therapeutic strategies in metabolic, neurodegenerative diseases and cancer.Mol Med. 2024 Mar 20;30(1):40. doi: 10.1186/s10020-024-00808-9. Mol Med. 2024. PMID: 38509524 Free PMC article. Review.
-
Proteostasis In The Endoplasmic Reticulum: Road to Cure.Cancers (Basel). 2019 Nov 14;11(11):1793. doi: 10.3390/cancers11111793. Cancers (Basel). 2019. PMID: 31739582 Free PMC article. Review.
-
Regulation of lipid metabolism by the unfolded protein response.J Cell Mol Med. 2021 Feb;25(3):1359-1370. doi: 10.1111/jcmm.16255. Epub 2021 Jan 4. J Cell Mol Med. 2021. PMID: 33398919 Free PMC article. Review.
Cited by
-
The Link Between Endoplasmic Reticulum Stress and Lysosomal Dysfunction Under Oxidative Stress in Cancer Cells.Biomolecules. 2025 Jun 25;15(7):930. doi: 10.3390/biom15070930. Biomolecules. 2025. PMID: 40723802 Free PMC article. Review.
References
Publication types
LinkOut - more resources
Full Text Sources

