Epidemiology and Risk Prediction Model of Multidrug-Resistant Organism Infections After Liver Transplant Recipients: A Single-Center Cohort Study
- PMID: 40281777
- PMCID: PMC12024721
- DOI: 10.3390/bioengineering12040417
Epidemiology and Risk Prediction Model of Multidrug-Resistant Organism Infections After Liver Transplant Recipients: A Single-Center Cohort Study
Abstract
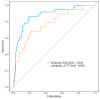
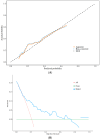
Objective: Accurate risk stratification at an early stage may reduce the incidence of infection and improve the survival rate of recipients by adopting targeted interventions. This study aimed to develop a nomogram to predict the risk of multidrug-resistant organism (MDRO) infections in liver transplant (LT) recipients. Methods: We retrospectively collected clinical data from 301 LT recipients and randomly divided them into a training set (210 cases) and validation set (91 cases) using a 7:3 split ratio. Factors related to the risk of MDRO infection after LT were determined using univariate and multivariate bidirectional stepwise logistic regression. The model's predictive performance and discrimination ability were evaluated using receiver operating characteristic (ROC) curves, calibration curves, and decision curve analysis (DCA). Results: 56 (18.60%) patients developed a MDRO infection, including 37 (17.62%) in the training cohort and 19 (20.88%) in the validation cohort. Ultimately, five factors related to MDRO infection after LT surgery were established: ascites (OR = 3.48, 95% CI [1.33-9.14], p = 0.011), total bilirubin (OR = 1.01, 95% CI [1.01-1.01], p < 0.001), albumin (OR = 0.85, 95% CI [0.75-0.96], p = 0.010), history of preoperative ICU stay (OR = 1.09, 95% CI [1.01-1.17], p = 0.009), and length of ICU stay (OR = 3.70, 95% CI [1.39-9.84], p = 0.019). The model demonstrated strong discrimination, and the area under the curve (AUC), sensitivity, and specificity of the training set were 0.88 (95% CI [0.81-0.94]), 0.82 (95% CI [0.76-0.87]), and 0.86 (95% CI [0.75-0.98]), respectively, while for the validation set, they were 0.77 (95% CI [0.65-0.90]), 0.76 (95% CI [0.67-0.86]), and 0.68 (95% CI [0.48-0.89]). The mean absolute error (MAE) in the validation cohort was 0.029, indicating a high accuracy. DCA showed a clinical benefit within a threshold probability range of 0.1 to 0.7. Conclusions: This study developed a clinically accessible nomogram to predict the risk of MDRO infection in LT recipients, enabling early risk stratification and the real-time assessment of infection risk based on the length of postoperative ICU stay. The model incorporates five easily obtainable clinical parameters (ascites, total bilirubin, albumin, preoperative ICU stay history, and length of ICU stay) and demonstrates strong predictive performance, facilitating the early identification of high-risk patients. Future research should focus on refining the model by incorporating additional clinical factors (e.g., immunosuppressive therapy adherence) and validating its generalizability in multicenter, large-sample cohorts to enhance its clinical utility.
Keywords: liver transplantation; multi-resistant organisms; nomogram; postoperative infection.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Oriol I., Sabé N., Simonetti A.F., Lladó L., Manonelles A., González J., Tubau F., Carratalà J. Changing trends in the aetiology, treatment and outcomes of bloodstream infection occurring in the first year after solid organ transplantation: A single-centre prospective cohort study. Transplant. Int. 2017;30:903–913. doi: 10.1111/tri.12984. - DOI - PubMed
-
- Viehman J.A., Clancy C.J., Clarke L., Shields R.K., Silveira F.P., Kwak E.J., Vergidis P., Hughes C., Humar A., Nguyen M.H. Surgical Site Infections After Liver Transplantation: Emergence of Multidrug-Resistant Bacteria and Implications for Prophylaxis and Treatment Strategies. Transplantation. 2016;100:2107–2114. doi: 10.1097/TP.0000000000001356. - DOI - PubMed
-
- Martin-Mateos R., Martínez-Arenas L., Carvalho-Gomes Á., Aceituno L., Cadahía V., Salcedo M., Arias A., Lorente S., Odriozola A., Zamora J., et al. Multidrug-resistant bacterial infections after liver transplantation: Prevalence, impact, and risk factors. J. Hepatol. 2024;80:904–912. doi: 10.1016/j.jhep.2024.02.023. - DOI - PubMed
LinkOut - more resources
Full Text Sources

