Genome-Wide Dissection of Novel QTLs and Genes Associated with Weed Competitiveness in Early-Backcross Selective Introgression-Breeding Populations of Rice (Oryza sativa L.)
- PMID: 40282278
- PMCID: PMC12025310
- DOI: 10.3390/biology14040413
Genome-Wide Dissection of Novel QTLs and Genes Associated with Weed Competitiveness in Early-Backcross Selective Introgression-Breeding Populations of Rice (Oryza sativa L.)
Abstract
The direct-seeded rice (DSR) system is poised to become the dominant rice cultivation method due to its advantages, including reduced water usage, less labor requirements, decreased greenhouse gas emissions, and improved adaptation to climate change. However, weeds, particularly jungle rice (Echinochloa colona), significantly hinder DSR and cause substantial yield losses. This study aimed to develop rice cultivars competitive against jungle rice through selective breeding, focusing on early seed germination (ESG) and seedling vigor (ESV). We utilized 181 early-backcross selective introgression breeding lines (EB-SILs) developed using Green Super Rice (GSR) technology by backcrossing Weed Tolerant Rice1 (WTR1) with three donor parents, Haoannong, Cheng Hui 448, and Y134. Using the tunable genotyping-by-sequencing (tGBS®, Data2Bio Technologies, Ames, IA, USA) method, we identified 3971 common single nucleotide polymorphisms (SNPs) that facilitated the mapping of 19 novel quantitative trait loci (QTLs) associated with weed competitiveness-eight linked to ESG traits and eleven to ESV traits. Notably, all QTLs were novel except qRPH1, linked to relative plant height at 14 and 21 days after sowing. Key QTLs were located on chromosomes 2, 3, 5, 6, 8, 9, 10, and 12. Candidate genes identified within these QTLs are implicated in the plant's response to various abiotic and biotic stresses. Our findings enhance the understanding of the genetic basis for ESG and ESV traits critical for weed competitiveness, supporting marker-assisted and genomic selection approaches for breeding improved rice varieties. Furthermore, this research lays the groundwork for employing gene expression, cloning, and CRISPR editing strategies to combat jungle rice, with potential applications for other weed species and contributing to effective integrated weed management in the DSR system.
Keywords: Green Super Rice; candidate genes; direct-seeded rice; jungle rice; quantitative trait loci; seedling vigor; single nucleotide polymorphisms; weed competitiveness.
Conflict of interest statement
The authors assert that this research review was carried out without affiliations with commercial or economic entities that could be interpreted as potential conflicts of interest. No conflicts of interest have been declared.
Figures

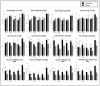

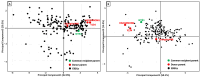


References
-
- Ali J., Anumalla M., Murugaiyan V., Li Z. Rice Improvement. Springer International Publishing; Cham, Switzerland: 2021. Green Super Rice (GSR) Traits: Breeding and Genetics for Multiple Biotic and Abiotic Stress Tolerance in Rice; pp. 59–97. - DOI
-
- Samal P., Babu S.C., Mondal B., Mishra S.N. The Global Rice Agriculture towards 2050: An Inter-Continental Perspective. Outlook Agric. 2022;51:164–172. doi: 10.1177/00307270221088338. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

