GC Content in Nuclear-Encoded Genes and Effective Number of Codons (ENC) Are Positively Correlated in AT-Rich Species and Negatively Correlated in GC-Rich Species
- PMID: 40282392
- PMCID: PMC12026676
- DOI: 10.3390/genes16040432
GC Content in Nuclear-Encoded Genes and Effective Number of Codons (ENC) Are Positively Correlated in AT-Rich Species and Negatively Correlated in GC-Rich Species
Abstract
Background/objectives: Codon usage bias affects gene expression and translation efficiency across species. The effective number of codons (ENC) and GC content influence codon preference, often displaying unimodal or bimodal distributions. This study investigates the correlation between ENC and GC rankings across species and how their relationship affects codon usage distributions.
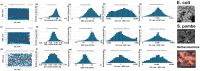
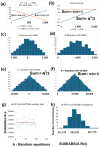
Methods: I analyzed nuclear-encoded genes from 17 species representing six kingdoms: one bacteria (Escherichia coli), three fungi (Saccharomyces cerevisiae, Neurospora crassa, and Schizosaccharomyces pombe), one archaea (Methanococcus aeolicus), three protists (Rickettsia hoogstraalii, Dictyostelium discoideum, and Plasmodium falciparum),), three plants (Musa acuminata, Oryza sativa, and Arabidopsis thaliana), and six animals (Anopheles gambiae, Apis mellifera, Polistes canadensis, Mus musculus, Homo sapiens, and Takifugu rubripes). Genes in all 17 species were ranked by GC content and ENC, and correlations were assessed. I examined how adding or subtracting these rankings influenced their overall distribution in a new method that I call Two-Rank Order Normalization or TRON. The equation, TRON = SUM(ABS((GC rank1:GC rankN) - (ENC rank1:ENC rankN))/(N2/3), where (GC rank1:GC rankN) is a rank-order series of GC rank, (ENC rank1:ENC rankN) is a rank-order series ENC rank, sorted by the rank-order series GC rank. The denominator of TRON, N2/3, is the normalization factor because it is the expected value of the sum of the absolute value of GC rank-ENC rank for all genes if GC rank and ENC rank are not correlated.
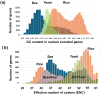
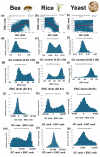
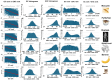
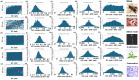
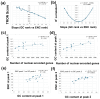
Results: ENC and GC rankings are positively correlated (i.e., ENC increases as GC increases) in AT-rich species such as honeybees (R2 = 0.60, slope = 0.78) and wasps (R2 = 0.52, slope = 0.72) and negatively correlated (i.e., ENC decreases as GC increases) in GC-rich species such as humans (R2 = 0.38, slope = -0.61) and rice (R2 = 0.59, slope = -0.77). Second, the GC rank-ENC rank distributions change from unimodal to bimodal as GC content increases in the 17 species. Third, the GC rank+ENC rank distributions change from bimodal to unimodal as GC content increases in the 17 species. Fourth, the slopes of the correlations (GC versus ENC) in all 17 species are negatively correlated with TRON (R2 = 0.98) (see Graphic Abstract).
Conclusions: The correlation between ENC rank and GC rank differs among species, shaping codon usage distributions in opposite ways depending on whether a species' nuclear-encoded genes are AT-rich or GC-rich. Understanding these patterns might provide insights into translation efficiency, epigenetics mediated by CpG DNA methylation, epitranscriptomics of RNA modifications, RNA secondary structures, evolutionary pressures, and potential applications in genetic engineering and biotechnology.
Keywords: CpG DNA methylation; GC content; bimodal distributions; codon bias; effective number of codons (ENC); epitranscriptomics; two-rank order normalization (TRON); unimodal distributions.
Conflict of interest statement
The author declares no conflict of interest.
Figures
Similar articles
-
Cross-species analysis of genic GC3 content and DNA methylation patterns.Genome Biol Evol. 2013;5(8):1443-56. doi: 10.1093/gbe/evt103. Genome Biol Evol. 2013. PMID: 23833164 Free PMC article.
-
[Synonymous codon usage bias in the rice cultivar 93-11 (Oryza sativa L. ssp. indica)].Yi Chuan Xue Bao. 2003 Apr;30(4):335-40. Yi Chuan Xue Bao. 2003. PMID: 12812058 Chinese.
-
Analysis of Codon Usage Bias in Chloroplast Genomes of Dryas octopetala var. asiatica (Rosaceae).Genes (Basel). 2024 Jul 9;15(7):899. doi: 10.3390/genes15070899. Genes (Basel). 2024. PMID: 39062678 Free PMC article.
-
Codon usage and codon pair patterns in non-grass monocot genomes.Ann Bot. 2017 Nov 28;120(6):893-909. doi: 10.1093/aob/mcx112. Ann Bot. 2017. PMID: 29155926 Free PMC article. Review.
-
Analysis of synonymous codon usage patterns in the edible fungus Volvariella volvacea.Biotechnol Appl Biochem. 2017 Mar;64(2):218-224. doi: 10.1002/bab.1538. Epub 2016 Dec 15. Biotechnol Appl Biochem. 2017. PMID: 27696508 Review.
Cited by
-
Comparative Analysis of Codon Usage Bias in Transcriptomes of Eight Species of Formicidae.Genes (Basel). 2025 Jun 27;16(7):749. doi: 10.3390/genes16070749. Genes (Basel). 2025. PMID: 40725406 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

