Computational Investigation of Montelukast and Its Structural Derivatives for Binding Affinity to Dopaminergic and Serotonergic Receptors: Insights from a Comprehensive Molecular Simulation
- PMID: 40283994
- PMCID: PMC12030116
- DOI: 10.3390/ph18040559
Computational Investigation of Montelukast and Its Structural Derivatives for Binding Affinity to Dopaminergic and Serotonergic Receptors: Insights from a Comprehensive Molecular Simulation
Abstract
Background/Objectives: Montelukast (MLK), a leukotriene receptor antagonist, has been associated with neuropsychiatric side effects. This study aimed to rationally modify MLK's structure to reduce these risks by optimizing its interactions with dopamine D2 (DRD2) and serotonin 5-HT1A receptors using computational molecular simulation techniques. Methods: A library of MLK derivatives was designed and screened using structural similarity analysis, molecular docking, molecular dynamics (MD) simulations, MM/PBSA binding free energy calculations, and ADME-Tox predictions. Structural similarity analysis, based on Tanimoto coefficient fingerprinting, compared MLK derivatives to known neuropsychiatric drugs. Docking was performed to assess initial receptor binding, followed by 100 ns MD simulations to evaluate binding stability. MM/PBSA calculations quantified binding affinities, while ADME-Tox profiling predicted pharmacokinetic and toxicity risks. Results: Several MLK derivatives showed enhanced DRD2 and 5-HT1A binding. MLK_MOD-42 and MLK_MOD-43 emerged as the most promising candidates, exhibiting MM/PBSA binding free energies of -31.92 ± 2.54 kcal/mol and -27.37 ± 2.22 kcal/mol for DRD2 and -30.22 ± 2.29 kcal/mol and -28.19 ± 2.14 kcal/mol for 5-HT1A, respectively. Structural similarity analysis confirmed that these derivatives share key pharmacophoric features with atypical antipsychotics and anxiolytics. However, off-target interactions were not assessed, which may influence their overall safety profile. ADME-Tox analysis predicted improved oral bioavailability and lower neurotoxicity risks. Conclusions: MLK_MOD-42 and MLK_MOD-43 exhibit optimized receptor interactions and enhanced pharmacokinetics, suggesting potential neuropsychiatric applications. However, their safety and efficacy remain to be validated through in vitro and in vivo studies. Until such validation is performed, these derivatives should be considered as promising candidates with optimized receptor binding rather than confirmed safer alternatives.
Keywords: 5-HT1A; D2 dopamine; molecular docking; molecular dynamics; montelukast; pharmacophore modeling.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

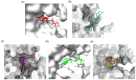


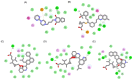
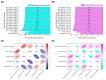



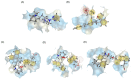
Similar articles
-
Exploring Radioiodinated Anastrozole and Epirubicin as AKT1-Targeted Radiopharmaceuticals in Breast Cancer: In Silico Analysis and Potential Therapeutic Effect with Functional Nuclear Imagining Implications.Molecules. 2024 Sep 4;29(17):4203. doi: 10.3390/molecules29174203. Molecules. 2024. PMID: 39275052 Free PMC article.
-
Structural modelling and in silico pharmacology of β-carboline alkaloids as potent 5-HT1A receptor antagonists and reuptake inhibitors.J Biomol Struct Dyn. 2023 Aug-Sep;41(13):6219-6235. doi: 10.1080/07391102.2022.2104376. Epub 2022 Jul 26. J Biomol Struct Dyn. 2023. PMID: 35881145
-
Unveiling Pharmacological Mechanisms of Bombyx mori (Abresham), a Traditional Arabic Unani Medicine for Ischemic Heart Disease: An Integrative Molecular Simulation Study.Pharmaceutics. 2025 Feb 24;17(3):295. doi: 10.3390/pharmaceutics17030295. Pharmaceutics. 2025. PMID: 40142959 Free PMC article.
-
Some Flavolignans as Potent Sars-Cov-2 Inhibitors via Molecular Docking, Molecular Dynamic Simulations and ADME Analysis.Curr Comput Aided Drug Des. 2022;18(5):337-346. doi: 10.2174/1573409918666220816113516. Curr Comput Aided Drug Des. 2022. PMID: 35975852
-
Discovery of New Dual-Target Agents Against PPAR-γ and α-Glucosidase Enzymes with Molecular Modeling Methods: Molecular Docking, Molecular Dynamic Simulations, and MM/PBSA Analysis.Protein J. 2024 Jun;43(3):577-591. doi: 10.1007/s10930-024-10196-y. Epub 2024 Apr 20. Protein J. 2024. PMID: 38642318
Cited by
-
From Lab to Clinic: How Artificial Intelligence (AI) Is Reshaping Drug Discovery Timelines and Industry Outcomes.Pharmaceuticals (Basel). 2025 Jun 30;18(7):981. doi: 10.3390/ph18070981. Pharmaceuticals (Basel). 2025. PMID: 40732273 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

