Chikungunya Replication and Infection Is Dependent upon and Alters Cellular Hexosylceramide Levels in Vero Cells
- PMID: 40284952
- PMCID: PMC12031450
- DOI: 10.3390/v17040509
Chikungunya Replication and Infection Is Dependent upon and Alters Cellular Hexosylceramide Levels in Vero Cells
Abstract
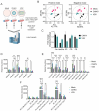
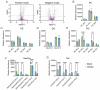
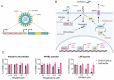
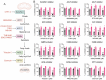
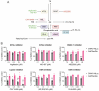
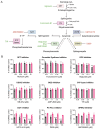
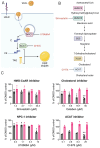
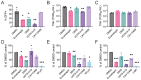
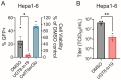
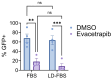
Chikungunya virus (CHIKV), a mosquito-borne alphavirus, causes significant global morbidity, including fever, rash, and persistent arthralgia. Utilizing untargeted lipidomics, we investigated how CHIKV infection alters host cell lipid metabolism in Vero cells. CHIKV infection induced marked catabolism of hexosylceramides, reducing their levels while increasing ceramide byproducts. Functional studies revealed a reliance on fatty acid synthesis, β-oxidation, and glycosphingolipid biosynthesis. Notably, inhibition of uridine diphosphate glycosyltransferase 8 (UGT8), essential for galactosylceramide production, significantly impaired CHIKV replication and entry in Vero cells. Sensitivity of CHIKV to UGT8 inhibition was reproduced in a disease-relevant cell line, mouse hepatocytes (Hepa1-6). CHIKV was also sensitive to evacetrapib, a cholesterol ester transfer protein (CETP) inhibitor, though the mechanism of inhibition appeared independent of CETP itself, suggesting an off-target effect. These findings highlight specific lipid pathways, particularly glycosphingolipid metabolism, as critical for CHIKV replication and further refine our understanding of how CHIKV exploits host lipid networks. This study provides new insights into CHIKV biology and suggests that targeted investigation of host lipid pathways may inform future therapeutic strategies.
Keywords: alphavirus; chikungunya virus; fatty acid synthesis; hexosylceramide; lipidomics.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures
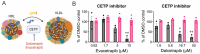
References
-
- Weaver S.C., Lecuit M. Chikungunya virus and the global spread of a mosquito-borne disease. N. Engl. J. Med. 2015;372:1231–1239. - PubMed
-
- Rodriguez-Morales A.J., Cardona-Ospina J.A., Fernanda Urbano-Garzon S., Sebastian Hurtado-Zapata J. Prevalence of Post-Chikungunya Infection Chronic Inflammatory Arthritis: A Systematic Review and Meta-Analysis. Arthritis Care Res. 2016;68:1849–1858. - PubMed
-
- Gogia A., B S., Kakar A. CHIKUNGUNYA: A Mortality Report. Open Forum Infect. Dis. 2017;4:S518.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

