Pharmacometabolomics Detects Various Unreported Metoprolol Metabolites in Urine of (Potential) Living Kidney Donors and Kidney Transplant Recipients
- PMID: 40285825
- PMCID: PMC12064448
- DOI: 10.1007/s40262-025-01502-7
Pharmacometabolomics Detects Various Unreported Metoprolol Metabolites in Urine of (Potential) Living Kidney Donors and Kidney Transplant Recipients
Abstract
Background and objective: Metoprolol is primarily metabolized via the polymorphic cytochrome P450-2D6 (CYP2D6) enzyme, which underlies interindividual variation in conversion rates and may benefit from pharmacogenetics-driven therapy personalization. However, the field relies heavily on knowledge of a drug's metabolism, often originating from early-phase clinical trials with single-dose administration in small samples of healthy volunteers. Pharmacogenetics could thus benefit from real-world drug metabolism studies.
Methods: We conducted a real-world drug metabolism study for metoprolol in 18 (potential) living kidney donors and 374 kidney transplant recipients from the Transplant Lines Food and Nutrition Biobank and Cohort Study (NCT02811835) using existing liquid chromatography-high resolution mass spectrometry pharmacometabolomic data.
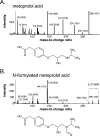
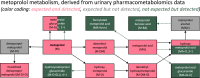
Results: In both groups, we confirmed the presence of seven expected metabolites, including the high-abundance substances metoprolol acid and hydroxymetoprolol. We were unable to detect deisopropylmetoprolol and a metabolite known as "H 119/68". However, we did find putative further oxidized forms, namely the expected variant of deisopropylmetoprolol in which the primary amine is removed and the leftover methyl group is oxidized into a carboxylic acid ("H 104/83") and an unknown/unreported metoprolol metabolite that we refer to as "metoprolol benzoic acid". Moreover, we found nine other previously unknown/unreported metabolites, putatively reflecting N-glucuronidated metoprolol, four glucuronidated versions of hydroxymetoprolol, and a formylated, a glucuronidated, and two hydroxylated versions of metoprolol acid. Interestingly, the same metabolites were detected in potential living kidney donors and kidney transplant recipients, and metabolite profiles did not differ between both groups in principal component analysis.
Conclusion: We found more metoprolol metabolites than previously reported, calling for replication studies and evaluation of pharmacogenetic testing approaches to realize safer, more effective metoprolol therapy.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Funding: Frank Klont gratefully acknowledges the funding provided by the Netherlands Organisation for Scientific Research NWO (domain Applied and Engineering Sciences) under Veni grant agreement no. 19060. The funder had no role in the study design, data collection, analysis, reporting, or the decision to submit for publication. Furthermore, the generation of the TransplantLines Food and Nutrition Biobank and Cohort Study (TxL-FN), trial registration number NCT02811835, was funded by the Top Institute Food and Nutrition, grant number A-1003. Continuation of the TransplantLines Food and Nutrition Biobank and Cohort Study was supported by grants from Astellas BV and Chiesi Pharmaceuticals BV and co-financed by the Dutch Ministry of Economic Affairs and Climate Policy by means of the public–private partnership allowance made available by Top Sector Life Sciences & Health to stimulate public–private partnerships. The funders had no role in the study design, data collection, analysis, reporting, or decision to submit for publication. Conflict of interest: The authors have no financial or proprietary interests in any material discussed in this article. Ethics approval: This study used existing data of KTRs and PLKD included in the TransplantLines Food and Nutrition Biobank and Cohort Study (NCT identifier NCT02811835), which was approved by the institutional review board of the University Medical Center Groningen (decision METc 2008/186) and adhered to the Declaration of Helsinki. Consent to participate: All participants provided written informed consent before study participation. Consent for publication: Not applicable. Availability of data: The pharmacometabolomic data used have been deposited in an open-access data repository, as can be found at: https://doi.org/10.26037/yareta:ybdgdynykfe6rkjjxb7d6oynoa [35]. Patient-level data for data on file in this manuscript are not available for sharing as there is a reasonable likelihood that individual patients could be re-identified. Further information can be requested from the authors. Author contributions: W.A.H., M.A.J.H., P.S., S.J.L.B., G.H., and F.K. wrote the manuscript. F.K. designed the research. W.A.H., M.A.J.H., and F.K. performed the research. W.A.H. and F.K. analyzed the data. S.J.L.B. and G.H. contributed new reagents/analytical tools.
Figures

Similar articles
-
Influence of CYP2D6*10 on the pharmacokinetics of metoprolol in healthy Korean volunteers.J Clin Pharm Ther. 2008 Oct;33(5):567-73. doi: 10.1111/j.1365-2710.2008.00945.x. J Clin Pharm Ther. 2008. PMID: 18834373 Clinical Trial.
-
Analysis of serum concentrations of metoprolol and its metabolite α-hydroxymetoprolol in patients with heart failure with reduced ejection fraction: a pilot study in routine health care.Expert Rev Clin Pharmacol. 2025 Jan-Feb;18(1-2):89-99. doi: 10.1080/17512433.2025.2450257. Epub 2025 Jan 16. Expert Rev Clin Pharmacol. 2025. PMID: 39757778
-
Metoprolol oxidation polymorphism in Brazilian elderly cardiac patients.J Pharm Pharmacol. 2013 Sep;65(9):1347-53. doi: 10.1111/jphp.12109. Epub 2013 Jul 21. J Pharm Pharmacol. 2013. PMID: 23927473
-
Simultaneous determination of metoprolol and its metabolites, α-hydroxymetoprolol and O-desmethylmetoprolol, in human plasma by liquid chromatography with tandem mass spectrometry: Application to the pharmacokinetics of metoprolol associated with CYP2D6 genotypes.J Sep Sci. 2014 Jun;37(11):1256-64. doi: 10.1002/jssc.201301353. Epub 2014 Apr 19. J Sep Sci. 2014. PMID: 24648255
-
Polymorphism of human cytochrome P450 2D6 and its clinical significance: Part I.Clin Pharmacokinet. 2009;48(11):689-723. doi: 10.2165/11318030-000000000-00000. Clin Pharmacokinet. 2009. PMID: 19817501 Review.
References
-
- Langreth R, Waldholz M. New era of personalized medicine: targeting drugs for each unique genetic profile. Oncologist. 1999;4:426–7. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

