Correlation of in vitro biofilm formation capacity with persistence of antibiotic-resistant Escherichia coli on gnotobiotic lamb's lettuce
- PMID: 40293242
- PMCID: PMC12093982
- DOI: 10.1128/aem.00299-25
Correlation of in vitro biofilm formation capacity with persistence of antibiotic-resistant Escherichia coli on gnotobiotic lamb's lettuce
Abstract
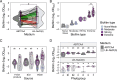
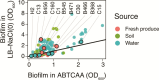
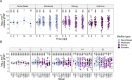
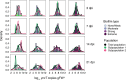
Bacterial contamination of fresh produce is a growing concern for food safety, as apart from human pathogens, antibiotic-resistant bacteria (ARB) can persist on fresh leafy produce. A prominent persistence trait in bacteria is biofilm formation, as it provides increased tolerance to stressful conditions. We screened a comprehensive collection of 174 antibiotic-susceptible and -resistant Escherichia coli originating from fresh leafy produce and its production environment. We tested the ability of these strains to produce biofilms, ranging from none or weak to extreme biofilm-forming bacteria. Next, we tested the ability of selected antibiotic-resistant isolates to colonize gnotobiotic lamb's lettuce (Valerianella locusta) plants. We hypothesized that a higher in vitro biofilm formation capacity correlates with increased colonization of gnotobiotic plant leaves. Despite a marked difference in the ability to form in vitro biofilms for a number of E. coli strains, in vitro biofilm formation was not associated with increased survival on gnotobiotic V. locusta leaf surfaces. However, all tested strains persisted for at least 21 days, highlighting potential food safety risks through unwanted ingestion of resistant bacteria. Population densities of biofilm-forming E. coli exhibited a complex pattern, with subpopulations more successful in colonizing gnotobiotic V. locusta leaves. These findings emphasize the complex behavior of ARB on leaf surfaces and their implications for human safety.IMPORTANCEEach raw food contains a collection of microorganisms, including bacteria. This is of special importance for fresh produce such as leafy salads or herbs, as these foods are usually consumed raw or after minimal processing, whereby higher loads of living bacteria are ingested than with a food that is heated before consumption. A common bacterial lifestyle involves living in large groups embedded in secreted protective substances. Such bacterial assemblies, so-called biofilms, confer high persistence and resistance of bacteria to external harsh conditions. In our research, we investigated whether stronger in vitro biofilm formation by antibiotic-resistant Escherichia coli correlates with better survival on lamb's lettuce leaves. Although no clear correlation was observed between biofilm formation capacity and population density on the salad, all tested isolates could survive for at least 3 weeks with no significant decline over time, highlighting a potential food safety risk independently of in vitro biofilm formation.
Keywords: Escherichia coli; antibiotic resistance; biofilms; fresh leafy produce; lamb's lettuce; plant-microbe interactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Influence of Plant Species, Tissue Type, and Temperature on the Capacity of Shiga-Toxigenic Escherichia coli To Colonize, Grow, and Be Internalized by Plants.Appl Environ Microbiol. 2019 May 16;85(11):e00123-19. doi: 10.1128/AEM.00123-19. Print 2019 Jun 1. Appl Environ Microbiol. 2019. PMID: 30902860 Free PMC article.
-
Adherence factors of enterohemorrhagic Escherichia coli O157:H7 strain Sakai influence its uptake into the roots of Valerianella locusta grown in soil.Food Microbiol. 2018 Dec;76:245-256. doi: 10.1016/j.fm.2018.05.016. Epub 2018 May 31. Food Microbiol. 2018. PMID: 30166148
-
Antibiotic resistance, virulence factors and biofilm formation ability in Escherichia coli strains isolated from chicken meat and wildlife in the Czech Republic.J Environ Sci Health B. 2017 Aug 3;52(8):570-576. doi: 10.1080/03601234.2017.1318637. Epub 2017 May 11. J Environ Sci Health B. 2017. PMID: 28494209
-
Life on the outside: role of biofilms in environmental persistence of Shiga-toxin producing Escherichia coli.Front Microbiol. 2014 Jul 1;5:317. doi: 10.3389/fmicb.2014.00317. eCollection 2014. Front Microbiol. 2014. PMID: 25071733 Free PMC article. Review.
-
Biofilm formation by enteric pathogens and its role in plant colonization and persistence.Microb Biotechnol. 2014 Nov;7(6):496-516. doi: 10.1111/1751-7915.12186. Microb Biotechnol. 2014. PMID: 25351039 Free PMC article. Review.
References
-
- Drissner D, Gekenidis M-T. 2023. Safety of food and beverages: fruits and vegetables. Reference Module in Food Science 1:10–19. doi:10.1016/B978-0-12-822521-9.00105-2 - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

