DNA damage in proximal tubules triggers systemic metabolic dysfunction through epigenetically altered macrophages
- PMID: 40295524
- PMCID: PMC12037803
- DOI: 10.1038/s41467-025-59297-x
DNA damage in proximal tubules triggers systemic metabolic dysfunction through epigenetically altered macrophages
Abstract
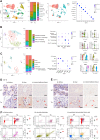
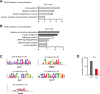
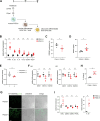
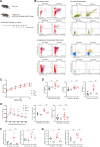
DNA damage repair is a critical physiological process closely linked to aging. The accumulation of DNA damage in renal proximal tubular epithelial cells (PTEC) is related to a decline in kidney function. Here, we report that DNA double-strand breaks in PTECs lead to systemic metabolic dysfunction, including weight loss, reduced fat mass, impaired glucose tolerance with mitochondrial dysfunction, and increased inflammation in adipose tissues and the liver. Single-cell RNA sequencing analysis reveals expansion of CD11c+ Ccr2+ macrophages in the kidney cortex, liver, and adipose tissues and Ly6Chi monocytes in peripheral blood. DNA damage in PTECs is associated with hypomethylation of macrophage activation genes, including Gasdermin D, in peripheral blood cells, which is linked to reduced DNA methylation at KLF9-binding motifs. Macrophage depletion ameliorates metabolic abnormalities. These findings highlight the impact of kidney DNA damage on systemic metabolic homeostasis, revealing a kidney-blood-metabolism axis mediated by epigenetic changes in macrophages.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures



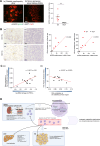
References
-
- Sedelnikova, O. A. et al. Senescing human cells and ageing mice accumulate DNA lesions with unrepairable double-strand breaks. Nat. Cell Biol.6, 168–170 (2004). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

