Replicative DNA polymerase epsilon and delta holoenzymes show wide-ranging inhibition at G-quadruplexes in the human genome
- PMID: 40298112
- PMCID: PMC12038398
- DOI: 10.1093/nar/gkaf352
Replicative DNA polymerase epsilon and delta holoenzymes show wide-ranging inhibition at G-quadruplexes in the human genome
Abstract
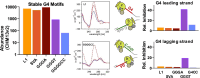
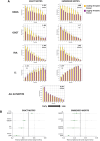
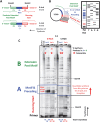
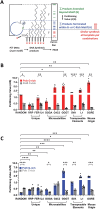
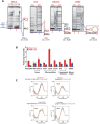
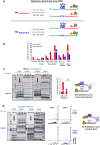
G-quadruplexes (G4s) are functional elements of the human genome, some of which inhibit DNA replication. We investigated replication of G4s within highly abundant microsatellite (GGGA, GGGT) and transposable element (L1 and SVA) sequences. We found that genome-wide, numerous motifs are located preferentially on the replication leading strand and the transcribed strand templates. We directly tested replicative polymerase ϵ and δ holoenzyme inhibition at these G4s, compared to low abundant motifs. For all G4s, DNA synthesis inhibition was higher on the G-rich than C-rich strand or control sequence. No single G4 was an absolute block for either holoenzyme; however, the inhibitory potential varied over an order of magnitude. Biophysical analyses showed the motifs form varying topologies, but replicative polymerase inhibition did not correlate with a specific G4 structure. Addition of the G4 stabilizer pyridostatin severely inhibited forward polymerase synthesis specifically on the G-rich strand, enhancing G/C strand asynchrony. Our results reveal that replicative polymerase inhibition at every G4 examined is distinct, causing complementary strand synthesis to become asynchronous, which could contribute to slowed fork elongation. Altogether, we provide critical information regarding how replicative eukaryotic holoenzymes navigate synthesis through G4s naturally occurring thousands of times in functional regions of the human genome.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nucleic Acids Research.
Conflict of interest statement
None declared.
Figures



Similar articles
-
DNA polymerase δ stalls on telomeric lagging strand templates independently from G-quadruplex formation.Nucleic Acids Res. 2013 Dec;41(22):10323-33. doi: 10.1093/nar/gkt813. Epub 2013 Sep 13. Nucleic Acids Res. 2013. PMID: 24038470 Free PMC article.
-
The in vitro fidelity of yeast DNA polymerase δ and polymerase ε holoenzymes during dinucleotide microsatellite DNA synthesis.DNA Repair (Amst). 2011 May 5;10(5):497-505. doi: 10.1016/j.dnarep.2011.02.003. Epub 2011 Mar 22. DNA Repair (Amst). 2011. PMID: 21429821 Free PMC article.
-
Complementary roles of Pif1 helicase and single stranded DNA binding proteins in stimulating DNA replication through G-quadruplexes.Nucleic Acids Res. 2019 Sep 19;47(16):8595-8605. doi: 10.1093/nar/gkz608. Nucleic Acids Res. 2019. PMID: 31340040 Free PMC article.
-
Guidelines for G-quadruplexes: I. In vitro characterization.Biochimie. 2023 Nov;214(Pt A):5-23. doi: 10.1016/j.biochi.2022.12.019. Epub 2022 Dec 31. Biochimie. 2023. PMID: 36596406 Review.
-
Structurally diverse G-quadruplexes as the noncanonical nucleic acid drug target for live cell imaging and antibacterial study.Chem Commun (Camb). 2023 Feb 2;59(11):1415-1433. doi: 10.1039/d2cc05945b. Chem Commun (Camb). 2023. PMID: 36636928 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

