Discovery of ATX968: An Orally Available Allosteric Inhibitor of DHX9
- PMID: 40298172
- PMCID: PMC12067447
- DOI: 10.1021/acs.jmedchem.5c00252
Discovery of ATX968: An Orally Available Allosteric Inhibitor of DHX9
Abstract
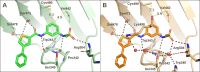
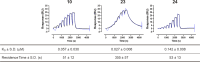
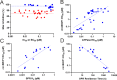
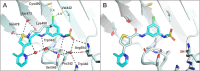
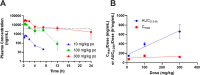
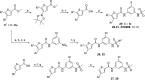
DHX9 is an RNA/DNA helicase integral in the maintenance of genome stability that has emerged as an attractive target for oncology drug discovery. Disclosed herein is the discovery and optimization of a series of DHX9 inhibitors. Compound 1 was identified as a partial inhibitor of DHX9 ATPase activity but a full inhibitor of unwinding activity. Binding of 1 to a pocket distinct from the ATP binding site was confirmed by X-ray crystallography, enabling structure-based drug optimization. During this optimization, a sulfur-halogen bond was identified that increased on-target residence time without impacting equilibrium binding affinity. Analysis shows that cell potency more closely correlates with residence time than with equilibrium measurements of binding affinity or biochemical potency. Further optimization of potency and ADME properties led to the identification of ATX968, a potent and selective DHX9 inhibitor that is efficacious in a tumor xenograft model of microsatellite instability-high (MSI-H) colorectal cancer.
Conflict of interest statement
The authors declare the following competing financial interest(s): All authors are or were employees of Accent Therapeutics while work was completed and may hold company stocks or stock options with Accent Therapeutics.
Figures
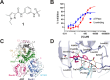


References
-
- Castro J.; Daniels M. H.; Brennan D.; Johnston B.; Gotur D.; Lee Y.-T.; Knockenhauer K. E.; Lu C.; Wu J.; Nayak S.; Collins C.; Bansal R.; Buker S. M.; Case A.; Liu J.; Yao S.; Sparling B. A.; Sickmier E. Allen.; Silver S. J.; Blakemore S. J.; Boriack-Sjodin P. A.; Duncan K. W.; Ribich S.; Copeland R. A. A Potent, Selective, Small-Molecule Inhibitor of DHX9 Abrogates Proliferation of Microsatellite Instable Cancers with Deficient Mismatch Repair. Cancer Res. 2025, 85, 758–776. 10.1158/0008-5472.CAN-24-0397. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

