GET3B is involved in chloroplast biogenesis and interacts with the thylakoidal ALB3 and ALB4 insertases
- PMID: 40299103
- PMCID: PMC12040988
- DOI: 10.1007/s00299-025-03500-2
GET3B is involved in chloroplast biogenesis and interacts with the thylakoidal ALB3 and ALB4 insertases
Abstract
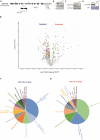
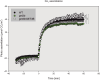
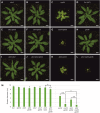
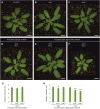
Proteomic, functional physiological analyses of get3b mutant plants highlight GET3B's role in chloroplast function. Genetic and interaction analyses indicate get3b and srp54 as mutual potentiators that might share terminal insertases. Protein targeting and insertion into membranes are essential for cellular organization and organelle function. The Guided Entry of Tail-anchored (GET) pathway facilitates the post-translational targeting and insertion of tail-anchored (TA) membrane proteins. Arabidopsis thaliana has four GET3 homologues, including AtGET3B and AtGET3D localized to chloroplasts. These photosynthetic organelles possess complex membrane systems, and the mechanisms underlying their protein targeting and membrane biogenesis are not fully understood. This study conducted a comprehensive proteomic analysis of get3b mutant plastids, which displayed significant alterations. Fluorometric based complex assembly as well as CO2 assimilation analyses confirmed that disruption of GET3B function displayed a significant impact on photosystem II assembly as well as carbon fixation, respectively, indicating a functional role in chloroplast biogenesis. Additionally, genetic interactions were found between GET3B and the two component STIC system, which cooperates with the cpSRP pathway which is involved in the co-translational sorting of thylakoid proteins. Further, physical interactions were observed between GET3B and the C-terminus of ALB3 and ALB4 in vitro and the full length proteins in vivo, indicating a role of GET3B in protein targeting and membrane integration within chloroplasts. These findings enhance our understanding of GET3B's involvement in stromal protein targeting and thylakoidal biogenesis.
Keywords: ALB3; ALB4; GET3B; STIC1; STIC2; cpSRP54.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interest: The authors have no relevant financial or non-financial interests to disclose. Ethical approval: The manuscript does not include data or description of human or animal patients.
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

