β-Catenin interacts with canonical RBPs including MSI2 to associate with a Wnt signalling mRNA network in myeloid leukaemia cells
- PMID: 40301545
- PMCID: PMC12256266
- DOI: 10.1038/s41388-025-03415-y
β-Catenin interacts with canonical RBPs including MSI2 to associate with a Wnt signalling mRNA network in myeloid leukaemia cells
Abstract
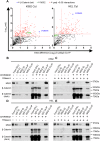
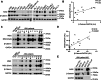
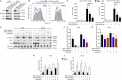
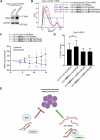
Wnt/β-catenin signalling is important for normal hematopoietic stem/progenitor cell (HSPC) biology and heavily implicated in acute and chronic myeloid leukaemia (AML and CML). The central mediator β-catenin is an attractive therapeutic target in myeloid neoplasms however its targeting has been hampered by a poor characterisation of its molecular interactions in haematopoietic cells, which will differ from its network in solid tissues. Our previous β-catenin interactome study identified the significant enrichment of RNA-binding proteins (RBP) implying post-transcriptional roles for β-catenin in myeloid cells. To identify β-catenin interacting mRNAs we performed β-catenin RNA-immunoprecipitation coupled to RNA-sequencing (RIP-seq) and identified significantly enriched Wnt signalling pathway transcripts. Using β-catenin cross-linking immunoprecipitation (CLIP) we demonstrated a limited capacity for β-catenin to bind RNA directly, implying dependence on other RBPs. β-Catenin was found to interact with Musashi-2 (MSI2) in both myeloid cell lines and primary AML patient samples, where expression was significantly correlated. MSI2 knockdown reduced Wnt signalling output (TCF/LEF activity), through suppression of LEF-1 expression and nuclear localisation. Through both RIP and CLIP we demonstrate MSI2 binds LEF1 mRNA in a partly β-catenin dependent fashion, and may impact the post-transcriptional control of LEF-1 expression. Finally, we show that MSI2-mediated expansion of human HSPCs could be partly driven through LEF1 regulation. This is the first study to experimentally demonstrate functional crosstalk between MSI2 and Wnt signalling in human cells, and indicates potential novel post-transcriptional roles for β-catenin in a haematological context.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
-
- Reya T, Duncan AW, Ailles L, Domen J, Scherer DC, Willert K, et al. A role for Wnt signalling in self-renewal of haematopoietic stem cells. Nature. 2003;423:409–14. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

