Establishment of a Triplex qPCR Assay for Differentiating Highly Virulent Genotype I Recombinant Virus From Low-Virulence Genotype I and Genotype II African Swine Fever Viruses Circulating in China
- PMID: 40303009
- PMCID: PMC12017118
- DOI: 10.1155/2024/6206857
Establishment of a Triplex qPCR Assay for Differentiating Highly Virulent Genotype I Recombinant Virus From Low-Virulence Genotype I and Genotype II African Swine Fever Viruses Circulating in China
Abstract
African swine fever virus (ASFV) poses serious threats to the global swine industry, food safety, and the economy. Since August 2018, different types of ASFVs have successively emerged in China, making ASF diagnostics more challenging. The highly virulent genotype I recombinant virus has gradually become the prevalent dominant strain and is identified by sequencing several of its genes, which is time-consuming and expensive. Here, we developed a triplex real-time quantitative PCR (qPCR) assay based on the ASFV B646L, X64R, and MGF_360-14L genes to differentiate highly virulent genotype I recombinant viruses from low-virulence genotype I and genotype II viruses in China. This method has high sensitivity and a limit of detection of 10 copies/reaction for standard plasmids, as well as good specificity without cross-reactions with the viral nucleic acids of porcine reproductive and respiratory syndrome virus (PRRSV), classical swine fever virus (CSFV), pseudorabies virus (PRV), porcine circovirus 2 (PCV 2), porcine circovirus 3 (PCV 3), porcine epidemic diarrhea virus (PEDV), transmissible gastroenteritis virus (TGEV), or porcine rotavirus (PoRV). Importantly, triplex qPCR can be used to quickly and accurately evaluate clinical samples and cell cultures infected with highly virulent genotype I virus, low-virulence genotype I virus, or genotype II virus. Thus, triplex qPCR provides an alternative tool for ASF surveillance in China.
Keywords: African swine fever virus; differential detection; genotype II; highly virulent genotype I; low virulent genotype I; triplex real-time quantitative PCR.
Copyright © 2024 Leilei Ding et al.
Conflict of interest statement
The authors declare no conflicts of interest.
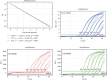
Figures



Similar articles
-
Development of a triplex real-time PCR assay for detection and differentiation of gene-deleted and wild-type African swine fever virus.J Virol Methods. 2020 Jun;280:113875. doi: 10.1016/j.jviromet.2020.113875. Epub 2020 Apr 23. J Virol Methods. 2020. PMID: 32333943
-
Development of a one-step multiplex qRT-PCR assay for the detection of African swine fever virus, classical swine fever virus and atypical porcine pestivirus.BMC Vet Res. 2022 Jan 18;18(1):43. doi: 10.1186/s12917-022-03144-4. BMC Vet Res. 2022. PMID: 35042532 Free PMC article.
-
Development of a multiplex qRT-PCR assay for detection of African swine fever virus, classical swine fever virus and porcine reproductive and respiratory syndrome virus.J Vet Sci. 2021 Nov;22(6):e87. doi: 10.4142/jvs.2021.22.e87. J Vet Sci. 2021. PMID: 34854269 Free PMC article.
-
Development and validation of a multiplex, real-time RT PCR assay for the simultaneous detection of classical and African swine fever viruses.PLoS One. 2013 Jul 26;8(7):e71019. doi: 10.1371/journal.pone.0071019. Print 2013. PLoS One. 2013. PMID: 23923045 Free PMC article.
-
Attenuated African swine fever viruses and the live vaccine candidates: a comprehensive review.Microbiol Spectr. 2024 Nov 5;12(11):e0319923. doi: 10.1128/spectrum.03199-23. Epub 2024 Oct 8. Microbiol Spectr. 2024. PMID: 39377589 Free PMC article. Review.
References
-
- Penrith M. L., Bastos A. D., Etter E. M. C., Beltrán-Alcrudo D. Epidemiology of African Swine Fever in Africa Today: Sylvatic Cycle Versus SocioEconomic Imperatives. Transboundary and Emerging Diseases . 2019;66(2):672–686. - PubMed
-
- WOAH. African Swine Fever (ASF)—Situation Report 46. 2024. https://www.woah.org/en/document/african-swine-fever-asf-situation-repor...
MeSH terms
LinkOut - more resources
Full Text Sources

