Molecular Detection of Pathogenic Leptospira sp. in Cetaceans from the Brazilian Coast
- PMID: 40303693
- PMCID: PMC12016789
- DOI: 10.1155/2023/7041089
Molecular Detection of Pathogenic Leptospira sp. in Cetaceans from the Brazilian Coast
Abstract
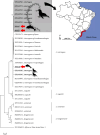
Leptospirosis is a zoonosis with ubiquitous distribution caused by spirochetes belonging to the genus Leptospira sp., endemic mainly in tropical and subtropical regions of the world and capable of infecting domestic animals, free-living animals, and humans. Although well documented in terrestrial animals and humans, little information is available on its distribution and impact on marine animals. There are few studies assessing cetaceans' health status, and even scarcer are those focused on leptospirosis research. In this context, considering the One Health approach, the present study aimed to investigate the occurrence of pathogenic Leptospira sp. in cetaceans on the Brazilian coast. Kidneys of 142 cetaceans belonging to 19 species were collected. DNA was extracted, and the diagnosis was performed by LipL32-polymerase chain reaction. Genetic characterization was conducted based on secY gene sequencing. Pathogenic Leptospira sp. DNA was detected in 14.8% (21/142) of the tested cetaceans, with coastal species presenting a significantly higher frequency (p-value = 0.03) of infected individuals (25%, 17/68) than oceanic species (7.5%, 4/53). It was possible to amplify and sequence three strains (one for Sotalia guianensis, one for Stenella clymene, and one for Pontoporia blainvillei), all of them identified as Leptospira interrogans, with high similarity with sequences from Icterohaemorrhagiae serogroup. Phylogenetic analysis revealed sequences from the present study grouped in species-specific unique clusters but very close to pinnipeds in the same area, evidencing the presence of two distinct haplotypes circulating on marine mammals in the region. We could demonstrate that cetaceans can act as carriers of pathogenic leptospires. Moreover, the proximity with anthropogenic areas could play an important role in leptospirosis' dynamics of transmission in a One Health context.
Copyright © 2023 Felipe D'Azeredo Torres et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest.
Figures
Similar articles
-
Pinnipeds carriers of pathogenic Leptospira: New data based on molecular characterization.Res Vet Sci. 2023 Feb;155:62-68. doi: 10.1016/j.rvsc.2022.12.012. Epub 2023 Jan 5. Res Vet Sci. 2023. PMID: 36634544
-
Genetic Evidence for a Potentially New Pathogenic Leptospira sp. Circulating in Bats from Brazilian Amazon.Transbound Emerg Dis. 2023 Sep 19;2023:9677047. doi: 10.1155/2023/9677047. eCollection 2023. Transbound Emerg Dis. 2023. PMID: 40303746 Free PMC article.
-
Genetic characteristics of pathogenic Leptospira in wild small animals and livestock in Jiangxi Province, China, 2002-2015.PLoS Negl Trop Dis. 2019 Jun 24;13(6):e0007513. doi: 10.1371/journal.pntd.0007513. eCollection 2019 Jun. PLoS Negl Trop Dis. 2019. PMID: 31233503 Free PMC article.
-
Didelphis albiventris as a carrier of Leptospira sp. in the central nervous tissue in the semiarid region of Northeast, Brazil.Comp Immunol Microbiol Infect Dis. 2020 Dec;73:101560. doi: 10.1016/j.cimid.2020.101560. Epub 2020 Oct 15. Comp Immunol Microbiol Infect Dis. 2020. PMID: 33099254 Review.
-
Leptospirosis in Unconventional Mammal Pets.Vet Sci. 2025 Mar 19;12(3):285. doi: 10.3390/vetsci12030285. Vet Sci. 2025. PMID: 40267017 Free PMC article. Review.
References
-
- Bossart G. D. Marine mammals as sentinel species for oceans and human health. Oceanography . 2006;19(2):134–137. - PubMed
-
- Aragones L. V., Roque M. A. A., Flores M. B., et al. The Philippine marine mammal strandings from 1998 to 2009: animals in the Philippines in peril? Aquatic Mammals . 2010;36(3):219–233. doi: 10.1578/AM.36.3.2010.219. - DOI
-
- Duignan P. J. Disease Investigations in Stranded Marine Mammals, 1999–2002 (DOC Science Internal Series 104) Wellington, NZ: Department of Conservation; 2003.
MeSH terms
LinkOut - more resources
Full Text Sources

