FleQ finetunes the expression of a subset of BrlR-activated genes to enable antibiotic tolerance by Pseudomonas aeruginosa biofilms
- PMID: 40304498
- PMCID: PMC12096822
- DOI: 10.1128/jb.00503-24
FleQ finetunes the expression of a subset of BrlR-activated genes to enable antibiotic tolerance by Pseudomonas aeruginosa biofilms
Abstract
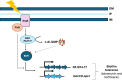
The transcriptional regulator FleQ contributes to Pseudomonas aeruginosa biofilm formation by activating the expression and biosynthesis of matrix exopolysaccharides in a manner dependent on c-di-GMP. However, little is known about the role of FleQ in the antibiotic tolerance phenotype of P. aeruginosa biofilms. Inactivation of fleQ impaired biofilm formation and rendered biofilms susceptible to tobramycin and norfloxacin. The phenotypes were similar to biofilms inactivated in sagS encoding the orphan sensor SagS that promotes the switch from planktonic to biofilm growth via BfiSR and antibiotic tolerance via BrlR. While FleQ was found to contribute to biofilm formation independently of SagS and BfiSR, FleQ instead converged with SagS-dependent regulation at the level of BrlR. This was supported by multicopy expression of sagS failing to restore biofilm antibiotic tolerance by ΔfleQ to wild-type levels (and vice versa) and by biofilms formed by the ΔfleQΔsagS double mutant being as susceptible as ΔfleQ and ΔsagS biofilms. Increased antibiotic susceptibility was independent of BrlR abundance or BrlR DNA binding but coincided with significantly reduced transcript abundance of the BrlR-activated mexCD-oprJ and PA1874-77, encoding an ABC transporter previously shown to contribute to the tolerance of biofilms to tobramycin and norfloxacin. FleQ- dependent regulation of gene expression was indirect. Co-immunoprecipitation and BACTH assays indicated FleQ to interact with SagS via its HisKA-Rec domain, likely suggesting FleQ and SagS to likely work in concert to enable biofilm antibiotic tolerance, by finetuning the expression of BrlR activated genes.IMPORTANCEIn P. aeruginosa, FleQ inversely regulates the expression of genes encoding flagella and biofilm matrix components, including exopolysaccharide (Pel, Psl) in a manner dependent on the levels of c-di-GMP. Our findings expand on the role of FleQ from regulating the transition to the biofilm mode of growth to FleQ contributing to the antimicrobial tolerance phenotype of biofilms, by FleQ affecting the expression of PA1874-77, a downstream target of the SagS-dependent transcriptional regulator BrlR. Importantly, our findings suggest FleQ works in concert with SagS, likely via FleQ-SagS protein-protein interactions, to enable the formation of inherently tolerant P. aeruginosa biofilms.
Keywords: BACTH; BrlR; DNA binding; biofilm-MBC; immunoblot; protein-protein interaction; pulldown; resistant to killing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Antimicrobial tolerance of Pseudomonas aeruginosa biofilms is activated during an early developmental stage and requires the two-component hybrid SagS.J Bacteriol. 2013 Nov;195(21):4975-87. doi: 10.1128/JB.00732-13. Epub 2013 Aug 30. J Bacteriol. 2013. PMID: 23995639 Free PMC article.
-
The ABC of Biofilm Drug Tolerance: the MerR-Like Regulator BrlR Is an Activator of ABC Transport Systems, with PA1874-77 Contributing to the Tolerance of Pseudomonas aeruginosa Biofilms to Tobramycin.Antimicrob Agents Chemother. 2018 Jan 25;62(2):e01981-17. doi: 10.1128/AAC.01981-17. Print 2018 Feb. Antimicrob Agents Chemother. 2018. PMID: 29180529 Free PMC article.
-
The Yin and Yang of SagS: Distinct Residues in the HmsP Domain of SagS Independently Regulate Biofilm Formation and Biofilm Drug Tolerance.mSphere. 2018 May 30;3(3):e00192-18. doi: 10.1128/mSphere.00192-18. Print 2018 Jun 27. mSphere. 2018. PMID: 29848761 Free PMC article.
-
Flip the switch: the role of FleQ in modulating the transition between the free-living and sessile mode of growth in Pseudomonas aeruginosa.J Bacteriol. 2024 Mar 21;206(3):e0036523. doi: 10.1128/jb.00365-23. Epub 2024 Mar 4. J Bacteriol. 2024. PMID: 38436566 Free PMC article. Review.
-
Recent perspectives on the molecular basis of biofilm formation by Pseudomonas aeruginosa and approaches for treatment and biofilm dispersal.Folia Microbiol (Praha). 2018 Jul;63(4):413-432. doi: 10.1007/s12223-018-0585-4. Epub 2018 Jan 19. Folia Microbiol (Praha). 2018. PMID: 29352409 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

