Role of Salivary MicroRNA as a Marker of Progesterone Resistance in Endometriosis: Preliminary Results from a Single-Institution Experience
- PMID: 40305232
- PMCID: PMC12025187
- DOI: 10.3390/biom15040493
Role of Salivary MicroRNA as a Marker of Progesterone Resistance in Endometriosis: Preliminary Results from a Single-Institution Experience
Abstract
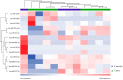
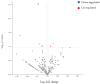
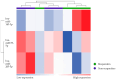
This feasibility study explores the potential of salivary microRNAs (miRNAs) as non-invasive biomarkers for diagnosing endometriosis and assessing treatment response. Almost one-third of patients with endometriosis do not respond to the standard progestin treatment due to various mechanisms of progesterone resistance. MiRNAs, recognized for their stability in body fluids and role in gene regulation, may offer new diagnostic and prognostic opportunities as they are involved in the pathogenic pathways of endometriosis and progesterone resistance. We sequenced salivary miRNAs in three cohorts of patients: control women without endometriosis and patients with endometriosis who responded and did not respond to standard progestin treatment. This aims to identify the differential miRNA expression profiles associated with therapeutic response to dienogest. The preliminary results demonstrate the feasibility of miRNA sequencing from saliva and reveal distinct miRNA profiles between responders, non-responders, and controls. Key miRNAs, including mir-3168, the mir-200a family, and mir-93-5p, emerged as potential biomarkers, showing significant differential expression linked to both endometriosis presence and treatment response. Further validation of these findings in larger cohorts could pave the way for miRNA-based diagnostic and prognostic tools, potentially reducing diagnostic delays and personalizing treatment approaches for endometriosis patients, also with new target therapies.
Keywords: biomarkers; endometriosis; miRNA; progesterone resistance; prognosis.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
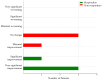

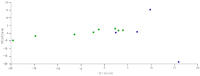


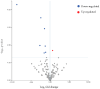



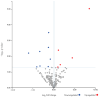
References
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical

