The motor neuron m6A repertoire governs neuronal homeostasis and FTO inhibition mitigates ALS symptom manifestation
- PMID: 40307231
- PMCID: PMC12043976
- DOI: 10.1038/s41467-025-59117-2
The motor neuron m6A repertoire governs neuronal homeostasis and FTO inhibition mitigates ALS symptom manifestation
Abstract
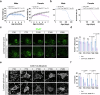
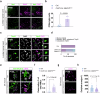
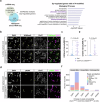
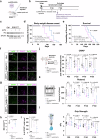
Amyotrophic lateral sclerosis (ALS) is a swiftly progressive and fatal neurodegenerative ailment marked by the degenerative motor neurons (MNs). Why MNs are specifically susceptible in predominantly sporadic cases remains enigmatic. Here, we demonstrated N6-methyladenosine (m6A), an RNA modification catalyzed by the METTL3/METTL14 methyltransferase complex, as a pivotal contributor to ALS pathogenesis. By conditional knockout Mettl14 in murine MNs, we recapitulate almost the full spectrum of ALS disease characteristics. Mechanistically, pervasive m6A hypomethylation triggers dysregulated expression of high-risk genes associated with ALS and an unforeseen reduction of chromatin accessibility in MNs. Additionally, we observed diminished m6A levels in induced pluripotent stem cell derived MNs (iPSC~MNs) from familial and sporadic ALS patients. Restoring m6A equilibrium via a small molecule or gene therapy significantly preserves MNs from degeneration and mitigates motor impairments in ALS iPSC~MNs and murine models. Our study presents a substantial stride towards identifying pioneering efficacious ALS therapies via RNA modifications.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. J.-A.C. and Y.-P.Y. declare a PCT and a US patent application related to this work.
Figures






References
-
- He, C. Grand challenge commentary: RNA epigenetics? Nat. Chem. Biol.6, 863 (2010). - PubMed
-
- Li, X., Xiong, X. & Yi, C. Epitranscriptome sequencing technologies: decoding RNA modifications. Nat. Methods14, 23 https://www.nature.com/articles/nmeth.4110#supplementary-information (2016). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

