Distinct gut microbiome characteristics and dynamics in patients with Parkinson's disease based on the presence of premotor rapid-eye movement sleep behavior disorders
- PMID: 40307949
- PMCID: PMC12042535
- DOI: 10.1186/s40168-025-02095-w
Distinct gut microbiome characteristics and dynamics in patients with Parkinson's disease based on the presence of premotor rapid-eye movement sleep behavior disorders
Abstract
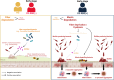
Background: Alpha-synuclein aggregation, a hallmark of Parkinson's disease (PD), is hypothesized to often begin in the enteric or peripheral nervous system in "body-first" PD and progresses through the vagus nerve to the brain, therefore REM sleep behavior disorder (RBD) precedes the PD diagnosis. In contrast, "brain-first" PD begins in the central nervous system. Evidence that gut microbiome imbalances observed in PD and idiopathic RBD exhibit similar trends supports body-first and brain-first hypothesis and highlights the role of microbiota in PD pathogenesis. However, further investigation is needed to understand distinct microbiome changes in body-first versus brain-first PD over the disease progression.
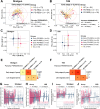
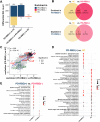
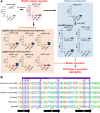
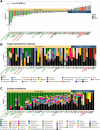
Results: Our investigation involved 104 patients with PD and 85 of their spouses as healthy controls (HC), with 57 patients (54.8%) categorized as PD-RBD(+) and 47 patients (45.2%) as PD-RBD(-) based on RBD presence before the PD diagnosis. We evaluated the microbiome differences between these groups over the disease progression through taxonomic and functional differential abundance analyses and carbohydrate-active enzyme (CAZyme) profiles based on metagenome-assembled genomes. The PD-RBD(+) gut microbiome showed a relatively stable microbiome composition irrespective of disease stage. In contrast, PD-RBD(-) microbiome exhibited a relatively dynamic microbiome change as the disease progressed. In early-stage PD-RBD(+), Escherichia and Akkermansia, associated with pathogenic biofilm formation and host mucin degradation, respectively, were enriched, which was supported by functional analysis. We discovered that genes of the UDP-GlcNAc synthesis/recycling pathway negatively correlated with biofilm formation; this finding was further validated in a separate cohort. Furthermore, fiber intake-associated taxa were decreased in early-stage PD-RBD(+) and the biased mucin-degrading capacity of CAZyme compared to fiber degradation.
Conclusion: We determined that the gut microbiome dynamics in patients with PD according to the disease progression depend on the presence of premotor RBD. Notably, early-stage PD-RBD(+) demonstrated distinct gut microbial characteristics, potentially contributing to exacerbation of PD pathophysiology. This outcome may contribute to the development of new therapeutic strategies targeting the gut microbiome in PD. Video Abstract.
Keywords: Biofilm; Carbohydrate-active enzymes; Microbiome; Parkinson’s disease; Rapid eye movement sleep behavior disorders.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: This study was approved by the Asan Medical Center Institutional Review Board (2019–0929) and was performed in accordance with the relevant guidelines and regulations, including the Declaration of Helsinki. All participants provided written informed consent at study enrollment. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures


Similar articles
-
The nasal and gut microbiome in Parkinson's disease and idiopathic rapid eye movement sleep behavior disorder.Mov Disord. 2018 Jan;33(1):88-98. doi: 10.1002/mds.27105. Epub 2017 Aug 26. Mov Disord. 2018. PMID: 28843021 Free PMC article.
-
Difference in gut microbial dysbiotic patterns between body-first and brain-first Parkinson's disease.Neurobiol Dis. 2024 Oct 15;201:106655. doi: 10.1016/j.nbd.2024.106655. Epub 2024 Aug 30. Neurobiol Dis. 2024. PMID: 39218360
-
[The gut microbiome in Parkinson's disease].Nervenarzt. 2019 Feb;90(2):160-166. doi: 10.1007/s00115-018-0601-6. Nervenarzt. 2019. PMID: 30171304 Review. German.
-
Functional implications of microbial and viral gut metagenome changes in early stage L-DOPA-naïve Parkinson's disease patients.Genome Med. 2017 Apr 28;9(1):39. doi: 10.1186/s13073-017-0428-y. Genome Med. 2017. PMID: 28449715 Free PMC article.
-
Research Progress on the Relationship between Parkinson's Disease and REM Sleep Behavior Disorder.J Integr Neurosci. 2024 Sep 14;23(9):166. doi: 10.31083/j.jin2309166. J Integr Neurosci. 2024. PMID: 39344226 Review.
Cited by
-
Genomic insights and metabolic profiling of gut commensal Luoshenia tenuis at strain level.NPJ Biofilms Microbiomes. 2025 Aug 5;11(1):153. doi: 10.1038/s41522-025-00793-9. NPJ Biofilms Microbiomes. 2025. PMID: 40764512 Free PMC article.
References
-
- Goedert M, Spillantini MG, Del Tredici K, Braak H. 100 years of Lewy pathology. Nat Rev Neurol. 2013;9:13–24. - PubMed
-
- Braak H, Del Tredici K, Rüb U, de Vos RA, Jansen Steur EN, Braak E. Staging of brain pathology related to sporadic Parkinson’s disease. Neurobiol Aging. 2003;24:197–211. - PubMed
-
- Stokholm MG, Danielsen EH, Hamilton-Dutoit SJ, Borghammer P. Pathological α-synuclein in gastrointestinal tissues from prodromal Parkinson disease patients. Ann Neurol. 2016;79:940–9. - PubMed
-
- Horsager J, Andersen KB, Knudsen K, Skjærbæk C, Fedorova TD, Okkels N, Schaeffer E, Bonkat SK, Geday J, Otto M, et al. Brain-first versus body-first Parkinson’s disease: a multimodal imaging case-control study. Brain. 2020;143:3077–88. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

