Design, synthesis, in silico studies, and apoptotic antiproliferative activity of novel thiazole-2-acetamide derivatives as tubulin polymerization inhibitors
- PMID: 40308265
- PMCID: PMC12040969
- DOI: 10.3389/fchem.2025.1565699
Design, synthesis, in silico studies, and apoptotic antiproliferative activity of novel thiazole-2-acetamide derivatives as tubulin polymerization inhibitors
Abstract
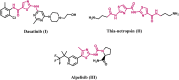
Introduction: Tubulin polymerization inhibitors have emerged as interesting anticancer therapies. We present the design, synthesis, and structural elucidation of novel thiazole-based derivatives to identify novel tubulin inhibitors with potent antiproliferative efficacy and strong inhibition of tubulin polymerization.
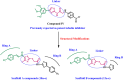
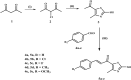
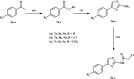
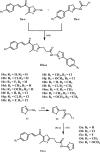
Methods: The novel compounds consist of two scaffolds. Scaffold A compounds 10a-e and scaffold B compounds 13a-e. the structures of the newly synthesized compounds 10a-e and 13a-e were validated using 1H NMR, 13C NMR, and elemental analysis.
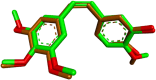
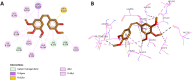
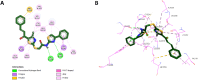
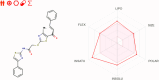
Results and discussion: The most effective antitubulin derivative was 10a, exhibiting an IC50 value of 2.69 μM. Subsequently, 10o and 13d exhibited IC50 values of 3.62 μM and 3.68 μM, respectively. These compounds exhibited more potency than the reference combretastatin A-4, which displayed an IC50 value of 8.33 μM. These compounds had no cytotoxic effects on normal cells, preserving over 85% cell viability at 50 μM. The antiproliferative experiment demonstrated that compounds 10a, 10o, and 13d displayed significant activity against four cancer cell lines, with average GI50 values of 6, 7, and 8 μM, equivalent to the reference's doxorubicin and sorafenib. Compounds 10a, 10o, and 13d were demonstrated to activate caspases 3, 9, and Bax, while down-regulating the anti-apoptotic protein Bcl2. Molecular docking studies demonstrated superior binding affinities for 10a (-7.3 kcal/mol) at the colchicine binding site of tubulin, forming key hydrophobic and hydrogen bonding interactions that enhance its activity. ADMET analysis confirmed favorable drug-like properties, establishing these compounds as promising candidates for further development as anticancer agents targeting tubulin polymerization.
Keywords: CA-4; antiproliferative; cell viability; colchicine; docking; tubulin.
Copyright © 2025 Al-Wahaibi, Elshamsy, Ali, Youssif, Bräse, Abdel-Aziz and El-Koussi.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
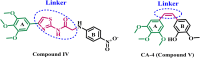

Similar articles
-
Design, Synthesis, in silico and in vitro Evaluation of New Combretastatin A-4 Analogs as Antimitotic Antitumor Agents.Med Chem. 2023;19(10):1018-1036. doi: 10.2174/1573406419666230530155741. Med Chem. 2023. PMID: 37259214
-
Synthesis and Molecular Docking Study of New Thiazole Derivatives as Potential Tubulin Polymerization Inhibitors.ACS Omega. 2022 Sep 7;7(37):33599-33613. doi: 10.1021/acsomega.2c05077. eCollection 2022 Sep 20. ACS Omega. 2022. PMID: 36157722 Free PMC article.
-
Synthesis of New Thiazole-Privileged Chalcones as Tubulin Polymerization Inhibitors with Potential Anticancer Activities.Pharmaceuticals (Basel). 2024 Aug 31;17(9):1154. doi: 10.3390/ph17091154. Pharmaceuticals (Basel). 2024. PMID: 39338317 Free PMC article.
-
Synthesis and Preclinical Evaluation of Indole Triazole Conjugates as Microtubule Targeting Agents that are Effective against MCF-7 Breast Cancer Cell Lines.Anticancer Agents Med Chem. 2021;21(8):1047-1055. doi: 10.2174/1871520620666200925102940. Anticancer Agents Med Chem. 2021. PMID: 32981511
-
Amide linked chalcone derivatives, a promising class of compounds with versatile biological effects.RSC Adv. 2025 Jun 5;15(24):19043-19068. doi: 10.1039/d5ra00834d. eCollection 2025 Jun 4. RSC Adv. 2025. PMID: 40476228 Free PMC article. Review.
Cited by
-
A novel thiazole-sulfonamide hybrid molecule as a promising dual tubulin/carbonic anhydrase IX inhibitor with anticancer activity.Front Chem. 2025 Jun 26;13:1606848. doi: 10.3389/fchem.2025.1606848. eCollection 2025. Front Chem. 2025. PMID: 40667533 Free PMC article.
References
-
- Abdelbaset M. S., Abdelrahman M. H., Bukhari S. N. A., Gouda A. M., Youssif B. G., Abdel-Aziz M., et al. (2021). Design, synthesis, and biological evaluation of new series of pyrrol-2 (3H)-one and pyridazin-3 (2H)-one derivatives as tubulin polymerization inhibitors. Bioorg. Chem. 107, 104522. 10.1016/j.bioorg.2020.104522 - DOI - PubMed
-
- Abdelbaset M. S., Abuo-Rahma G.E.-D. A., Abdelrahman M. H., Ramadan M., Youssif B. G., Bukhari S. N. A., et al. (2018). Novel pyrrol-2 (3H)-ones and pyridazin-3 (2H)-ones carrying quinoline scaffold as anti-proliferative tubulin polymerization inhibitors. Bioorg. Chem. 80, 151–163. 10.1016/j.bioorg.2018.06.003 - DOI - PubMed
-
- Abo-Ashour M. F., Eldehna W. M., George R. F., Abdel-Aziz M. M., Elaasser M. M., Gawad N. M. A., et al. (2018). Novel indole-thiazolidinone conjugates: design, synthesis and whole-cell phenotypic evaluation as a novel class of antimicrobial agents. Eur. J. Med. Chem. 160, 49–60. 10.1016/j.ejmech.2018.10.008 - DOI - PubMed
-
- Afzal O., Ali A., Ali A., Altamimi A. S. A., Alossaimi M. A., Bakht M. A., et al. (2023). Synthesis and anticancer evaluation of 4-Chloro-2-((5-aryl-1, 3, 4-oxadiazol-2-yl) amino) phenol analogues: an insight into experimental and theoretical studies. Molecules 28 (16), 6086. 10.3390/molecules28166086 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

