SREBP2 as a central player in cancer progression: potential for targeted therapeutics
- PMID: 40308757
- PMCID: PMC12041066
- DOI: 10.3389/fphar.2025.1535691
SREBP2 as a central player in cancer progression: potential for targeted therapeutics
Abstract
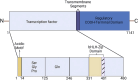
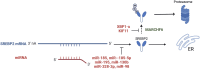
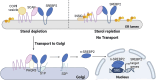
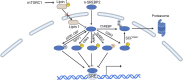
Recent studies have identified the reprogramming of lipid metabolism as a critical hallmark of malignancy. Enhanced cholesterol uptake and increased cholesterol biosynthesis significantly contribute to the rapid growth of tumors, with cholesterol also playing essential roles in cellular signaling pathways. Targeting cholesterol metabolism has emerged as a promising therapeutic strategy in oncology. The sterol regulatory element-binding protein-2 (SREBP2) serves as a primary transcriptional regulator of genes involved in cholesterol biosynthesis and is crucial for maintaining cholesterol homeostasis. Numerous studies have reported the upregulation of SREBP2 across various cancers, facilitating tumor progression. This review aims to provide a comprehensive overview of the structure, biological functions, and regulatory mechanisms of SREBP2. Furthermore, we summarize that SREBP2 plays a crucial role in various cancers and tumor microenvironment primarily by regulating cholesterol, as well as through several non-cholesterol pathways. We also particularly emphasize therapeutic agents targeting SREBP2 that are currently under investigation. This review seeks to enhance our understanding of SREBP2's involvement in cancer and provide theoretical references for cancer therapies that target SREBP2.
Keywords: SREBP2; cancer; cancer therapy; cholesterol metabolism; tumor microenvironment.
Copyright © 2025 Chen, Chen, Li, Yu, Lin, Wen, Zhang, Chen, Yi, Zhong and Li.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Investigation of Potential Drug Targets for Cholesterol Regulation to Treat Alzheimer's Disease.Int J Environ Res Public Health. 2023 Jun 24;20(13):6217. doi: 10.3390/ijerph20136217. Int J Environ Res Public Health. 2023. PMID: 37444065 Free PMC article.
-
Cholesterol Metabolism in CNS Diseases: The Potential of SREBP2 and LXR as Therapeutic Targets.Mol Neurobiol. 2025 May;62(5):6283-6307. doi: 10.1007/s12035-024-04672-w. Epub 2025 Jan 7. Mol Neurobiol. 2025. PMID: 39775479 Review.
-
The RORɣ/SREBP2 pathway is a master regulator of cholesterol metabolism and serves as potential therapeutic target in t(4;11) leukemia.Oncogene. 2024 Jan;43(4):281-293. doi: 10.1038/s41388-023-02903-3. Epub 2023 Nov 29. Oncogene. 2024. PMID: 38030791 Free PMC article.
-
Sterol-responsive element-binding protein (SREBP) 2 down-regulates ATP-binding cassette transporter A1 in vascular endothelial cells: a novel role of SREBP in regulating cholesterol metabolism.J Biol Chem. 2004 Nov 19;279(47):48801-7. doi: 10.1074/jbc.M407817200. Epub 2004 Sep 8. J Biol Chem. 2004. PMID: 15358760
-
Cholesterol Metabolic Reprogramming in Cancer and Its Pharmacological Modulation as Therapeutic Strategy.Front Oncol. 2021 May 24;11:682911. doi: 10.3389/fonc.2021.682911. eCollection 2021. Front Oncol. 2021. PMID: 34109128 Free PMC article. Review.
References
Publication types
LinkOut - more resources
Full Text Sources

