Targeting PCNA/PARP1 axis inhibits the malignant progression of hepatocellular carcinoma
- PMID: 40313621
- PMCID: PMC12043649
- DOI: 10.3389/fphar.2025.1571786
Targeting PCNA/PARP1 axis inhibits the malignant progression of hepatocellular carcinoma
Abstract
Introduction: Proliferating cell nuclear antigen (PCNA) is associated with the proliferation and recurrence of various cancers, and its high expression is associated with poor prognosis in hepatocellular carcinoma (HCC) patients. However, the mechanistic role of PCNA in HCC progression remains poorly understood. This study aimed to investigate how PCNA regulates DNA damage repair and cell cycle progression in HCC, with a focus on its interaction with poly (ADP-ribose) polymerase 1 (PARP1) and therapeutic implications.
Methods: PCNA was targeted genetically and pharmacologically in HCC cells to assess its effects on DNA damage repair and cell cycle arrest. Protein-protein interactions between PCNA and PARP1 were validated through co-immunoprecipitation and functional assays. The sensitivity of HCC cells to the PARP1 inhibitor Olaparib was evaluated under PCNA inhibition. Synergistic effects of AOH1160 (a PCNA inhibitor) and Olaparib were tested in vitro and in vivo using proliferation assays, DNA damage quantification, and cell cycle analysis. Prognostic relevance of PCNA expression was analyzed using TCGA datasets.
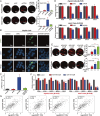
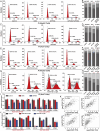
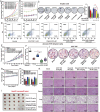
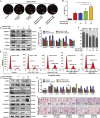
Results: Targeting PCNA suppressed DNA damage repair and induced cell cycle arrest in HCC cells. Mechanistically, PARP1 was identified as a downstream target of PCNA and directly interacted with PCNA. Inhibiting the expression or activity of PCNA increased the sensitivity of HCC cells to the PARP1 inhibitor, Olaparib. In addition, AOH1160 and Olaparib synergistically inhibited the proliferation, DNA damage repair and cell cycle progression of HCC cells. Elevated PCNA levels correlated with unfavorable HCC prognosis, supporting its role as a therapeutic biomarker. In vivo experiments also confirmed that repression of the PCNA/PARP1 axis significantly reduced HCC tumor growth.
Discussion: This study elucidates the relationship between PCNA and PARP1 in regulating the malignant progression of HCC, and highlight the pivotal role of PCNA/PARP1 axis in DNA damage repair and cell cycle progression. The correlation between elevated PCNA levels and unfavorable prognosis underscores its potential as a therapeutic biomarker. Repression of PCNA/PARP1 axis significantly inhibits the malignant proliferation of HCC cells both in vitro and in vivo. Collectively, the study provides a mechanistic foundation for therapies targeting PCNA/PARP1 axis.
Keywords: DNA damage repair; PARP1; PCNA; cell cycle progression; hepatocellular carcinoma.
Copyright © 2025 Li, Yong, Chen, Zeng, Zhang, Wang and Zhang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




References
LinkOut - more resources
Full Text Sources
Miscellaneous

