The modifying effect of mutant LRRK2 on mutant GBA1-associated Parkinson disease
- PMID: 40315377
- PMCID: PMC12228092
- DOI: 10.1093/hmg/ddaf062
The modifying effect of mutant LRRK2 on mutant GBA1-associated Parkinson disease
Erratum in
-
Correction to: The modifying effect of mutant LRRK2 on mutant GBA1-associated Parkinson disease.Hum Mol Genet. 2025 Jul 20;34(15):1341. doi: 10.1093/hmg/ddaf112. Hum Mol Genet. 2025. PMID: 40554637 Free PMC article. No abstract available.
Abstract
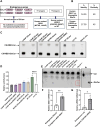
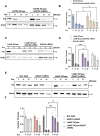
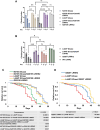
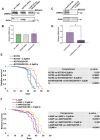
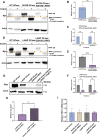
Parkinson disease (PD) is the second most common neurodegenerative disease. While most cases are sporadic, in ~ 5%-10% of PD patients the disease is caused by mutations in several genes, among them GBA1 (glucocerebrosidase beta 1) and LRRK2 (leucine-rich repeat kinase 2), both prevalent among the Ashkenazi Jewish population. LRRK2-associated PD tends to be milder than GBA1-associated PD. Several recent clinical studies have suggested that carriers of both GBA1 and LRRK2 mutations develop milder PD compared to that observed among GBA1 carriers. These findings strongly suggested an interplay between the two genes in the development and progression of PD. In the present study Drosophila was employed as a model to investigate the impact of mutations in the LRRK2 gene on mutant GBA1-associated PD. Our results strongly indicated that flies expressing both mutant genes exhibited milder parkinsonian signs compared to the disease developed in flies expressing only a GBA1 mutation. This was corroborated by a decrease in the ER stress response, increase in the number of dopaminergic cells, elevated levels of tyrosine hydroxylase, reduced neuroinflammation, improved locomotion and extended survival. Furthermore, a significant decrease in the steady-state levels of mutant GBA1-encoded GCase was observed in the presence of mutant LRRK2, strongly implying a role for mutant LRRK2 in degradation of mutant GCase.
Keywords: Drosophila; GBA1; LRRK2; Parkinson disease.
© The Author(s) 2025. Published by Oxford University Press.
Figures




Similar articles
-
[Autophagy Impairment in Parkinson's Disease: Approaches to Therapy].Mol Biol (Mosk). 2025 Jan-Feb;59(1):60-79. Mol Biol (Mosk). 2025. PMID: 40542632 Review. Russian.
-
P2RX7, an adaptive immune response gene, is associated with Parkinson's disease risk and age at onset.J Parkinsons Dis. 2024 Nov;14(8):1575-1583. doi: 10.1177/1877718X241296015. Epub 2024 Dec 1. J Parkinsons Dis. 2024. PMID: 39957192
-
Lysosomal glucocerebrosidase is needed for ciliary Hedgehog signaling: A convergent pathway contributing to Parkinson's disease.Proc Natl Acad Sci U S A. 2025 Aug 5;122(31):e2504774122. doi: 10.1073/pnas.2504774122. Epub 2025 Jul 30. Proc Natl Acad Sci U S A. 2025. PMID: 40737317 Free PMC article.
-
Genetic testing for Parkinson's disease in Israel: Insights from the Rostock Parkinson's Disease (ROPAD) study.Parkinsonism Relat Disord. 2025 Aug;137:107940. doi: 10.1016/j.parkreldis.2025.107940. Epub 2025 Jun 28. Parkinsonism Relat Disord. 2025. PMID: 40617169
-
LRRK2-mediated mitochondrial dysfunction in Parkinson's disease.Biochem J. 2025 May 28;482(11):721-39. doi: 10.1042/BCJ20253062. Biochem J. 2025. PMID: 40440058 Free PMC article. Review.
References
-
- Poewe W, Seppi K, Tanner CM. et al. Parkinson disease. Nat Rev Dis Primer 2017;3:17013. - PubMed
-
- Dauer W, Przedborski S. Parkinson’s disease: mechanisms and models. Neuron 2003;39:889–909. - PubMed
-
- Jankovic J. Parkinson’s disease: clinical features and diagnosis. J Neurol Neurosurg Psychiatry 2008;79:368–376. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

