Analysis of stress response in multiple bacterial pathogens using a network biology approach
- PMID: 40316612
- PMCID: PMC12048639
- DOI: 10.1038/s41598-025-91269-5
Analysis of stress response in multiple bacterial pathogens using a network biology approach
Abstract
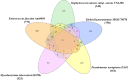
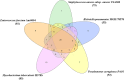
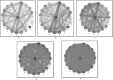
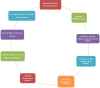
Stress response in bacterial pathogens promotes adaptation, virulence and antibiotic resistance. In this study, a network approach is applied to identify the common central mediators of stress response in five emerging opportunistic pathogens; Enterococcus faecium Aus0004, Staphylococcus aureus subsp. aureus USA300, Klebsiella pneumoniae MGH 78,578, Pseudomonas aeruginosa PAO1, and Mycobacterium tuberculosis H37Rv. A Protein-protein interaction network (PPIN) was constructed for each stressor using Cytoscape3.7.1 from the differentially expressed genes obtained from Gene expression omnibus datasets. A merged PPIN was constructed for each bacterium. Hub-bottlenecks in each network were the central stress response proteins and common pathways enriched in stress response were identified using KOBAS3.0. 31 hub-bottlenecks were common to each individual stress response, merged networks in all five pathogens and an independent cross stress (CS) response dataset of Escherichia coli. The 31 central nodes are in the RpoS mediated general stress regulon and also regulated by other stress response systems. Analysis of the 20 common metabolic pathways modulating stress response in all five bacteria showed that carbon metabolism pathway had the highest crosstalk with other pathways like amino acid biosynthesis and purine metabolism pathways. The central proteins identified can serve as targets for novel wide-spectrum antibiotics to overcome multidrug resistance.
Keywords: Bacterial pathogen; Centrality; Differentially expressed genes; Protein protein interaction network; Stress response.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

