Hijacking the MDM2 E3 Ligase with Novel BRD4-Targeting Proteolysis-Targeting Chimeras in Pancreatic Cancer Cells
- PMID: 40317844
- PMCID: PMC12247027
- DOI: 10.1002/cbic.202500133
Hijacking the MDM2 E3 Ligase with Novel BRD4-Targeting Proteolysis-Targeting Chimeras in Pancreatic Cancer Cells
Abstract
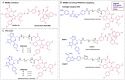
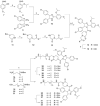
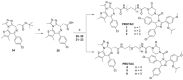
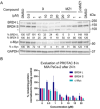
The phenotypic effect induced by a Proteolysis-Targeting Chimera (PROTAC) can depend on several factors, including the E3 ligase recruited. For the discovery of a first-in-class PROTAC for a target of interest, the E3 ligases commonly hijacked remain the Von Hippel-Lindau (VHL) and Cereblon (CRBN) since potent and accessible ligands are readily available to recruit them. Mouse double minute 2 (MDM2) E3 ligase stands out because it regulates p53 levels to maintain cellular homeostasis. However, the synthesis of the most potent MDM2 ligands remains very complex. Herein, the discovery of novel MDM2-recruiting PROTACs incorporating rac-Nutlin-3 as a ligand with an easier synthetic tractability is reported, further demonstrating its potential in this technology. The most promising degrader, PROTAC 3, showed preferential degradation of the BRD4 short isoform (BRD4 S) and c-Myc compared with MZ1, a validated VHL-based PROTAC.
Keywords: BRD4; HDM2; MDM2; MIA PaCa‐2; PROTACs; pancreatic cancer; targeted protein degradation.
© 2025 The Author(s). ChemBioChem published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Brown D. G., Wobst H. J., J. Med. Chem. 2021, 64, 2312. - PubMed
-
- Vetma V., O’Connor S., Ciulli A., Annu. Rev. Cancer Biol. 2024,. 9, 119.
-
- Bashore C., Prakash S., Johnson M. C., Conrad R. J., Kekessie I. A., Scales S. J., Ishisoko N., Kleinheinz T., Liu P. S., Popovych N., Wecksler A. T., Zhou L., Tam C., Zilberleyb I., Srinivasan R., Blake R. A., Song A., Staben S. T., Zhang Y., Arnott D., Fairbrother W. J., Foster S. A., Wertz I. E., Ciferri C., Dueber E. C., Nat. Chem. Biol. 2023, 19, 55. - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

