Synergistic bioinformatics and sophisticated machine learning unveil ferroptosis-driven regulatory pathways and immunotherapy potential in breast carcinoma
- PMID: 40320501
- PMCID: PMC12050258
- DOI: 10.1007/s12672-025-02393-7
Synergistic bioinformatics and sophisticated machine learning unveil ferroptosis-driven regulatory pathways and immunotherapy potential in breast carcinoma
Abstract
Background: The intersection of aberrant iron metabolism and the rapidly advancing field of immunotherapy has emerged as a critical focus in breast cancer (BRCA) therapeutics. Ferroptosis, a distinct form of iron-dependent cell death driven by lipid peroxidation, has garnered increasing attention for its pivotal role in cancer progression.
Methods: Utilizing extensive datasets from TCGA and GEO, this research extracted a wealth of biological data, including mRNA splicing indices, genomic aberrations, copy number variations (CNV), tumor mutational burden (TMB), and diverse clinical information. Through precise Lasso regression analysis, this research constructed a prognostic model that elucidates the molecular interactions of FRGs in BRCA. Concurrent co-expression network analyses were performed to explore the dynamic interplay between gene expression patterns and FRGs, revealing potential regulatory mechanisms.
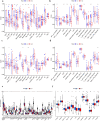
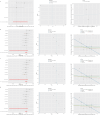
Results: This research analysis revealed significant overexpression of FRGs in high-risk BRCA samples, highlighting their prognostic relevance beyond traditional clinical parameters. GSVA identified immune response and cancer-related pathways as predominantly active in high-risk groups, suggesting ferroptosis as a central modulator within the tumor microenvironment. Notably, genes such as ACTL8, VGF, and CPLX2 emerged as markers of tumorigenesis, while IL33 and TP63 were identified as potential key regulators of cancer progression, each exhibiting distinct expression profiles across risk levels. Furthermore, this research incorporated gene correlations, CNV profiles, SNP arrays, and drug susceptibility analyses, contributing to the advancement of precision oncology.
Conclusions: The integration of bioinformatics and machine learning in this study underscores a strong correlation between FRG expression patterns and BRCA prognosis, affirming their potential as precise biomarkers for personalized immunotherapy.
Keywords: BRCA; CNV; Drug prediction; Drug prediction; Ferroptosis; Immune checkpoint; Immunity; m6a.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participation: This manuscript is not a clinical trial, hence the ethics approval and consent to participation is not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures






References
LinkOut - more resources
Full Text Sources
