Engineering Assembloids to Mimic Graft-Host Skeletal Muscle Interaction
- PMID: 40320876
- PMCID: PMC12232128
- DOI: 10.1002/adhm.202404111
Engineering Assembloids to Mimic Graft-Host Skeletal Muscle Interaction
Abstract
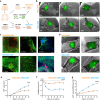
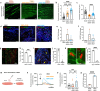
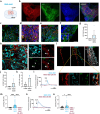
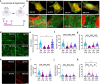
Skeletal muscle (SkM) tissue engineering aims to generate in vitro 3D products that can be implanted in patients to replace or repair damaged muscles. Having a humanized in vitro model able to mimic the interaction between the innervated recipient and the engineered SkMs at a functional level would greatly help in the evaluation of the graft potential. Here, a 3D in vitro model is developed that allows to investigation of the function, stability, and adaptability of the human neuromuscular (NM) system in response to an engineered SkM construct. To achieve this, decellularized SkMs (dSkM)-based constructs are used as engineered SkM and human neuromuscular organoids (NMOs) as the recipient-like NM system to create graft-host SkM assembloids. We observed the migration of myogenic cells and invasion of neural axons from the NMO to the engineered SkM construct in the assembloids, with the generation of functional neuromuscular junctions (NMJs). Finally, assembloids are able to regenerate following acute damage, with SkM regeneration and functional recovery. Despite being limited by the absence of immunocompetent cells and vasculature, the data showed that the assembloid represents a useful tool to evaluate in vitro the response of the human innervated SkM to a potential tissue-engineered SkM graft.
Keywords: assembloids; muscle regeneration; muscle stem cell; neuromuscular organoids; organoids; skeletal muscle; tissue engineering.
© 2025 The Author(s). Advanced Healthcare Materials published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
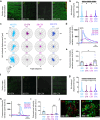
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

