Characterization and identification of extrachromosomal circular DNA in cholangiocarcinoma
- PMID: 40323971
- PMCID: PMC12052172
- DOI: 10.1371/journal.pone.0322173
Characterization and identification of extrachromosomal circular DNA in cholangiocarcinoma
Abstract
Extrachromosomal circular DNAs (eccDNAs) have gained attention as key players in cancer heterogeneity, potentially associated with elevated oncogene copy numbers in many cancers. While the presence of eccDNA in both normal and cancer cells is confirmed, its influence on gene-level alterations in cancer cells remains largely unexplored. This study delves into the genomic profiles of eccDNA in cholangiocarcinoma (CCA), an aggressive biliary tract cancer with extensive heterogeneity and diverse molecular alterations, using a modified long-read CircleSeq method. We reveal distinct eccDNA characteristics in CCA compared to non-tumor cells, focusing on genic components and chromosomal origins. Analysing read depth differences in oncogene-containing eccDNA; we identified potential eccDNA candidates that may be relevant for CCA biology. Subsequent bioinformatics analysis was performed using the established CReSIL tool, revealing distinct patterns of these oncogenes, particularly genes in the RAS/BRAF pathway, suggesting a potential functional role. These findings highlight the remarkable heterogeneity and diverse origins of eccDNA in CCA. This study establishes the first profiling of eccDNA in cholangiocarcinoma and paves the way for further investigation of its potential contribution to oncogene amplification and disease progression.
Copyright: © 2025 Win et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

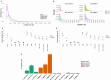
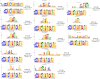
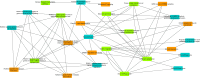

References
-
- Banales JM, Cardinale V, Carpino G, Marzioni M, Andersen JB, Invernizzi P, et al. Expert consensus document: Cholangiocarcinoma: current knowledge and future perspectives consensus statement from the European Network for the Study of Cholangiocarcinoma (ENS-CCA). Nature Reviews Gastroenterology and Hepatology. 2016;13:261–80. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

