Structure and nucleic acid interactions of the SΔ60 domain of the hepatitis delta virus small antigen
- PMID: 40324079
- PMCID: PMC12088457
- DOI: 10.1073/pnas.2411890122
Structure and nucleic acid interactions of the SΔ60 domain of the hepatitis delta virus small antigen
Abstract
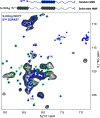
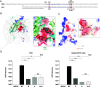
Infection with hepatitis delta virus (HDV) causes the most severe form of viral hepatitis, affecting more than 15 million people worldwide. HDV is a small RNA satellite virus of the hepatitis B virus (HBV) that relies on the HBV envelope for viral particle assembly. The only specific HDV component is the ribonucleoprotein (RNP), which consists of viral RNA (vRNA) associated with the small (S) and large (L) delta antigens (HDAg). While the structure of the HDAg N-terminal assembly domain is known, here we address the structure of the remaining SΔ60 protein using NMR. We show that SΔ60 contains two intrinsically disordered regions separated by a helix-loop-helix motif and that this structure is conserved in the full-length protein. Solution NMR analysis revealed that SΔ60 binds to both full-length and truncated vRNA, highlighting the role of the helical regions in submicromolar affinity interactions. The resulting complex contains approximately 120 SΔ60 proteins per RNA. Our results provide a model for the arginine-rich domains in RNP assembly and RNA interactions. In addition, we show that a cluster of acidic residues within the structured region of SΔ60 is critical for HDV replication, possibly mimicking the nucleosome acidic patch involved in the recruitment of chromatin remodelers. Our work thus provides the molecular basis for understanding the role of the C-terminal RNA-binding domain of S-HDAg in HDV infection.
Keywords: NMR; S-HDAg; acidic cluster; hepatitis delta virus; protein–RNA interaction.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures





References
-
- Hepojoki J., et al. , “Create one new realm (Ribozyviria) including one new family (Kolmioviridae) including genus Deltavirus and seven new genera for a total of 15 species” (Tech. Rep. 2020.012D, International Committee for Taxonomy of Viruses proposal (Taxoprop), 2020).
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

